- Asa
- 1980 1st AAC
- Contributed Papers
- Surface Management Of Arable Soils
- Tillage Components Of Conservation Cropping In The South Burnett
- Chemical Weed Control Under Stubble Mulching
- Edaphic Properties Of Fly Ash From Australian Power Stations
- Effect Of Different Tillage Systems On Wheat Production
- Effect Of Wheat Stubble Management During Fallow
- No Tillage Wheat Production In Northern New South Wales
- Runoff And Soil Loss From Contour Bays As Affected By Surface Conditions
- Effect Of Surface Cover On Infiltration Into A Self Mulching Soil
- The Use Of Polyvinyl Alcohol To Improve Emergence Of Vegetable Crops Grown On Fine Textured Goulburn Valley Soils
- The Implication Of High Flotation Applicators In Australian Agriculture
- The Effect Of Stubble Level On Runoff And Soil Loss Under Simulated Rainfall
- Soil Management In The Lockyer Valley
- Short Term Benefits Of Zero Tillage In Tropical Grain Production
- No Tillage For Conservation Or Arable Soils In The Northern Wheat Belt Of Australia
- Comparison Of Effects Of Direct Drilling And Ploughing On Cereal Production In The United Kingdom
- Structural Amelioration As A Factor In The Management Of Heavy Textured Soils
- Stubble Retention Study In Grain Sorghum In Central Queensland
- A Conservation Cropping System For The South Burnett
- Australian Pastures And Their Management
- The Survival Of Medic Seeds Following Ingestion Of Intact Pods By Sheep
- Herbage Attributes, Diet Preference And Pasture Management
- Herbage Biomass, Ponded Infiltration And Sulfur In A Disturbed Woodland
- Competitive Abilities Of Lucerne And Sorrel Growing In Acid Soils
- Use Of Ecological Benchmark Sites In Natural Pasture Condition Surveys In Australia
- Effect Of Defoliation And Clover Scorch On The Competitive Relations Of Two Subterranean Clover Cultivars In Mixture
- Pasture Establishment Using Oats As A Companion Crop On The Northern Tablelands Of N.S.W.
- Crop Sequences, Crop Pasture Rotations And Soil Fertility
- Nitrogen Economy Of Lupin Wheat Rotations
- Effect Of Lupin And Barley Rotations On Grain Yield And Soil Nitrogen
- Use Of Lucerne Leys For Maintaining The Productivity Of Wheat Growing Soils
- A Ley Farming System For The Semi Arid Tropics
- The Value Of A Legume Ley In Crop Rotations
- Oilseeds And Grain Legumes In The Southern Wheatbelt Of N.S.W.
- The Effects Of Lupin Management On Wheat Yields In Lupin Wheat Rotation
- A Pasture And Crop Sequence For Intensive Beef Cattle Production
- The Effect Of Tillage On The Fate Of Hard Seeds Of Subterranean Clover
- Nematodes A Major Factor In Crop Sequence Effects On Darling Downs Wheat.
- Nitrogen Economy In A Crop Sequence Study On The Darling Downs
- Effect Of Mung Beans On Soil Mineral Nitrogen On The Eastern Darling Downs.
- Genetic Exploitation Of The Environment
- Drought Resistance In Wheat
- Evaluating Wheat And Rapeseed Genotypes In Relation To Environment
- Phosphorus Utilization In Contrasting Pasture Legumes
- Overcoming Agronomic Shortcomings Of "opaque 2" Maize By Selection For Genetic Modifiers
- Yield Trends From Australian Wheats
- Cassava Cultivar Evaluation In South East Queensland
- Transference Of Sorghum Midge Resistance In To Agronomically Acceptable Lines
- A New Look At Guar In Central Queensland
- Performance Of Four Lupin Species In Central Queensland
- Irrigated Summer Forage Crops In Central Western N.S.W.
- Breeding Higher Yielding Lupins, Rapeseed, High Lysine Barley And Phalaris For South Eastern Australia
- The Significance Of Mediterranean Plant Introduction For Increasing The Winter Growth Of Pasture In The High Rainfall Areas Of Victoria
- Adaptation Of Cajanus Cajan (pigeon Pea) In Sub Tropical Australia
- Relative Resistance Of Snap Beans (phaseolus Vulgaris L) To Mechanical Injury Of Seed As Influenced By Maturation Temperature
- Vine Breeding At Merbein
- Breeding French Bean Cultivars For Winter Production
- The Role Of Varietal Control In The Protection Of Intensive Cropping Systems
- Insect, Disease And Weed Management
- The Prospects For Biocontrol Augmentation In Cotton And Soybean Systems
- Control Of Seed Harvesting Ants
- Insect Pest Management And The Queensland Cotton Industry
- Computer Retrieval Of Plant Pesticide Registration Information
- Survival Of Alternaria Carthami On Safflower Residue
- The Starwheel Sprayer A New Design For Foliage Spray Penetration Of Vegetables
- Effect Of Herbicides On Nitrophilous Weeds And The Resultant Establishment And Development Of Surface Sown Pasture Species
- Susceptibility Of Grain Sorghum Cultivars To 3,4 D And Picloram Plus 3,4 D
- The Potential Significance Of Plant Derived Chemicals In Crop/weed Associations
- Control Of Mexican Poppy (Argemone Mexicana Forma Ochroleuca) By Manipulation Of Seeding Rate
- Biological Control Of Weeds Modest Investments Can Give Large Returns
- Methomyl An Ovicide For Control Of Heliothis Spp. In A Cotton Pest Management System
- Land Resources And Horticultural Production
- Land Use And Catchment Management
- Problems Of Fertility Related To Long Periods Of Pasture Improvement And Management In A Small Urban Water Supply Catchment
- Rehabilitation Of Open Cut Mines In The Hunter Valley Of New South Wales
- Land Management Needs In The Bremer And Lockyer Catchments
- An Assessment Of Potential Land Use In Queensland
- Strip Cropping For Erosion Control On The Darling Downs
- Management Units For Agricultural Soils
- The Re Use Of Agricultural Land After Open Cut Coal Mining
- Land Use Studies In Western Queensland
- Land Use And Its Effect On Soil Conservation Catchment Management In North Western New South Wales
- Improved Plant Management
- The Essentials For Cotton Agroecosystem Optimisation: Current Problems And Future Prospects
- Yield Performance In The Australian Wheat Growing Industry
- Delayed Planting Of Short Season Soybeans In Narrow Rows And High Densities
- Some Limitations To High Soybean Yields In Southern N.S.W.
- Effect Of Plant Population Density On Grain Yield And Lodging Of Three Maize Cultivars
- Mepiquat Chloride A New Growth Regulant For Cotton
- Response Of Raingrown Soybeans To Rate And Placement Of Superphosphate On Low P Soils
- Direct Drilling Of Soybean Varieties In Coastal N.S.W.
- Modelling Soil Water For Direct Drilling And Conventional Cropping Strategies In Southern N.S.W.
- Yield Ceiling For Grain Sorghum In The Ord River Irrigation Area
- Time Of Harvest And Seed Quality In Lupins
- A Farm Test For Seed Quality In Lupins
- Improving Plant Management Through Modelling Plant/environment Interactions
- Methods Of Using Rongai Lablab For Beef Cattle Production
- Soybeans As A Dryland Crop In North West New South Wales
- Saturated Soil Culture Of Soybeans A New Agronomic System
- Spatial Arrangements And Wheat Yields
- Effects Of Spotted Alfalfa Aphids And Blue Green Aphids On The Dry Matter Production Of Some Lucerne Varieties
- Effect Of Sowing Rate On The Establishment Of Dryland Lucerne
- Response Of Cotton Cultivars To Planting Date
- Accelerated Ageing Tests Useful Measures Of Soybean Seed Planting Quality
- An Objective Basis For The Selection Of Minimum Legal Seed Germination Standards
- Simazine, A Potential Growth Regulator
- Evaluation Of Four Introduced Temperate Grass Species Under Grazing At Glen Innes, N.S.W.
- Yield Of Starch And Soluble Carbohydrates In Perennial Forage Sorghums
- Mixed Cropping For The Production Of Feed Rations
- Effect Of Nitrogen On The Development Of Yield Components In Wheat
- Developments In Fresh Market Tomato Production
- Blueberries In Australia
- Significant Areas Of Research In Sub Tropical Fruit Crops
- Rapid Propagation Techniques For Potato Cultivars
- Potential Utilisation Of Reclaimed Water For Vegetable Production
- Close Planting Improves Returns For Growing Apples
- Bean Seed Quality In Burdekin Crops: Effect Of Time And Method Of Harvest
- Processing Tomatoes For Machine Harvesting
- Glue Sprays For Anti Shattering Treatment Of Horticultural And Pasture Seed Crops
- Animal Production From Summer Forage Crops In Cool Environments
- Optimisation Of Plant Nutrition
- The Response Of Soybean Varieties To Zinc
- Nutrient Limitations On Yield And Quality Of Peanuts Grown In Four Soils From The Kingaroy Area I Phosphorus
- Nutrient Limitations On Yield And Quality Of Peanuts Grown In Four Soils From The Kingaroy Area II Molybdenum
- Nutrient Limitation On Yield And Quality Of Peanuts Grown In Four Soils In The Kingaroy Area III Sulphur
- The Fertilizer Requirements Of Continuous Wheat
- Nitrogen Availability And Wheat Performance Following Rice Crops
- Spatial And Temporal Variability Of Soil Test P Values Under Permanent Pasture
- Prediction Of Yield Response By Soybeans In The South Burnett, Queensland, Using Soil Tests
- Nutritional Investigations In Cotton, Emerald Irrigation Area
- Sulphur Status Of Pasture Legumes In The Semi Arid Wheatbelt
- The Effect Of Potassium On French Bean Seed Yield And Seed Vigour
- Nitrogen X Seeding Rate Interactions In Wheat In Northern N.S.W.
- Elemental Sulphur Fertilizer Use In The Northern Tablelands Of N.S.W.
- Prediction Of Lucerne Responses To Lime From Soil Tests
- Limitations To Efficient Use Of Nitrogen For Irrigated Crops
- Potential For Increased Efficiency In Crop And Animal Production By Applying Nutrients As A Foliar Spray
- Grain Yield And Protein Content In Wheat Cultivars In Response To Time Of Application Of Nitrogen Fertilizer
- The Residual Effects Of Fertilizer Zinc On A Black Earth Soil From North Western New South Wales
- Nitrogen Application Strategies For Rice
- Surface Management Of Arable Soils
- Conference Information
- Invited Papers
- Surface Management Of Arable Soils
- Australian Pastures And Their Management
- Crop Sequences, Crop Pasture Rotations And Soil Fertility
- Genetic Exploitation Of The Environment
- Insect, Disease And Weed Management
- Land Resources And Horticultural Production
- Land Use And Catchment Management
- Improved Plant Management
- Optimisation Of Plant Nutrition
- Contributed Papers
- 1982 2nd AAC
- Background Papers: Problems Facing Australian Agriculture
- Some Of The Problems Facing Agriculture In New South Wales
- Problems Facing Agriculture In Victoria
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Some Of The Problems Facing Agriculture In New South Wales
- Concluding Review
- Contributed Papers
- Agricultural Systems
- New Medic Pasture Systems To Increase Production In Pasture/cereal Rotations In The Victorian Mallee
- Supplementing Native Pastures With Leucaliva Llucoclpiala Cv. Peru (leucaena) In An Extensive Beef Cattle Production System
- Conservation Grazing Management In The Sub Humid Region Of Central Queensland
- Liveweight Gain Of Lambs On Lucerne, Other Legumes And Perennial Grass Sub Clover Pasture Over Summer
- Balancing Pasture Feed Supply With Animal Demands During Pasture Development And In Time Of Drought
- Changes In Agricultural Systems Of Southern Australia
- Changing Agricultural Systems And Insect Pests In South Australia
- Dryland Cropping In The Western Division Of New South Wales
- The Control Of Take All In The Victorian Mallee
- Improved Soil Management In NE Victoria
- Evidence For Disease Control As A Factor In Improved Wheat Yield In A Lupin Wheat Rotation
- The Effects Of Land And Fertilizer Management Techniques On Rice Productivity Under Intensive Rotations
- Chemical Control Of Silverleaf Nightshade Using A Ropewick Applicator
- Development Of Controlled Release Formulations For Agricultural Use In Australia
- Evaluation Of Weed Control Strategies A Simulation Study Of Nassella Trichotoma, Serrated Tussock
- The Effects Of Soil Compaction On The Growth Of Wheat And Subterranean Clover
- Conservation Management Of Pineapple Lands In Southeast Queensland
- Botanal A Versatile Field And Computing Package For Assessing Yield And Botanical Composition In Grazing Experiments
- Horticulture
- The Effects Of Seed Soaking, Temperature, And Other Environmental Factors On The Germination Of Jojoba (simmondsia Ciiinensis)
- Chemical Regulation Of Fruit Loosening As An Aid To Mechanical Harvesting In Fruit Crops
- Reducing Premature Nut Drop In Macadamia With N.A.A. Sprays
- Alternative Intensive Production Systems For Stonefruit And Citrus Crops
- Tea District Establishment And Yield Trials In Queensland
- Controlling Seed Borne Fungi By The Use Of Heat And Salt
- Irrigation And Water Use
- Irrigation Of Linseed At Emerald (B) Response To Irrigation In The Field
- Irrigation Of Linseed At Emerald (C) Economic Optimum Irrigation Regime
- The Need For Improved Irrigation Management To Attain High Yields Of Wheat In A Semi Arid Environment
- Fodder Production From Irrigated Crops In Northern Victoria
- Productivity Of Irrigated Pastures In Northern Victoria
- Potential Productivity Of Irrigated Perennial Pasture
- The Low Productivity Of Irrigated Agriculture Is Irrigation Method A Key Factor?
- Landforming In Northern Victoria Are Our Soils Suited To This Technology?
- Timing Pasture Irrigation With A "home Made" Evaporation Tank
- 308 The Lower Limit Of Extractable Soil Water For Crops Grown On A Cracking Clay Soil.
- The Effects Of Transient Waterlogging On The Yield Of Sunflower And Sorghum
- Wet Soil Culture Of Soybeans: Plant Growth And Yield
- Irrigation Of Sunflowers In The Southeast Of South Australia
- Comparison Of Water Use By Sunflower At Two Row Spacings
- Comparison Of Furrow And Sprinkler Irrigation Systems For Grain Sorghum Production In The Ord Irrigation Area
- Response Of Cassava To Irrigation
- The Effect Of Nitrogen Fertilizer On The Response Of The Potato Crop To Irrigation
- Irrigation Of Linseed At Emerald (a) Response To Moisture Stress
- Nutrition
- Lime For Cereals On Acid Soils In Northern New South Wales
- Responses To Lime By Cereals
- Effect Of Liming Acid Krasnozems On The Yield Of Poppy (Papaver Somniferum L.)
- Root Growth In Soils Rendered Acid By Improved Pastures
- Soil Management Studies And Growth Of Subterranean Clover In North East Victoria
- Long Term Trends Associated With Usf Of Superphosphate On Pasture, I. Yields Of Clover
- Long Term Trends Associated With Use Of Superphosphate On Pasture. Ii. Spatial Distribution Of Pasture Yield And Composition.
- Long Term Trends Associated With Superphosphate Use On Pasture, III. Response Of Clover To Potassium After Prolonged Treatment.
- Partially Acidulated Rock Phosphates (parp) As Fertiliser For Perennial Pastures
- Phosphate Research At Wagga Wagga
- Fungi For More Efficient Use Of P Fertiliser
- Leaf Acid Phosphatase And The Phosphorus Status Of Wheat
- Availability Of Phosphate Fertilizer Residues In A Soil Of Western Queensland During Three Years Of Cropping
- Phosphorus Application Strategies For Rice (Starbonnet) In The Lower Burdekin Area, Queensland
- The Response Of Cassava To Phosphorus Fertilizer On Five Soils In South East Queensland
- Diammonium Phosphate As A Source Of N And P For Sorghum In The South Burnett
- The Effect Of Phosphorus Availability And Soil Type On Manganese Accumulation In Lupinus Albus L.
- Foliar Manganese For Lupins
- Phosphorus And Sulfur Interactions In Soil Plant Systems. I. Uptake Of P And S By Plants Grown In Undrained Pasture Cores.
- Phosphorus And Sulfur Interactions In Soil Plant Systems. II. Field Studies On The Effect Of Phosphorus On Soil Available Sulfur In Uncrazed Plots.
- Phosphorus And Sulfur Interactions In Soil Plant Systems, III. Effects Of Grazing And Superphosphate On Soil Sulfur Profiles
- Available Sulphur Levels In Mallee Soils
- Fertilizing Field Crops Pre Plant Banding Of Dual Applications Of Nitrogen And Phosphorus Fertilizers
- The Effect Of Nitrogen And Phosphorus Application On Yield Of Raingrown Summer Crops In Central Queensland.1. Sorghum In Dawson And Callide Valleys
- The Effect Of Nitrogen And Phosphorus Application On Yield Of Raingrown Summer Crops In Central Queensland. 2. Sorghum And Sunflower Central Highlands
- A Simple Model For Predicting Nitrogen Fertilizer Requirement Of Cereal Crops
- Field Studies On The Fate Of Urea Applied To Flooded Rice
- Effect Of Changing Nitrogen Supply On Growth, Nitrogen Accumulation And Seed Characteristics Of Sunflower
- The Availability Of Early Applied Nitrogen For Winter Crops In Queensland
- Alternative Sources Of Nitrogen For Wheat On Solonised Brown Soils In The Victorian Mallee
- Nitrogen Recovery And Grain Yield Of Inga Rice
- Increasing Efficiency Of Water Run Fertilization
- Nitrogen Fixation In Grain Legumes
- The Utilization Of Wheat Straw As An Energy Source For Biological Nitrogen Fixation
- Nitrate Reduction And Dinitrogen Fixation In Chickpea
- A Comparison Of Three Inoculation Treatments On Lupins
- Field Evaluation Of Rh/zob/um Strains On Sainfoin (onobrychis Viciifolia Scop.)
- The Effects Of Defoliation, Flower Removal, Applied Nitrogen, And Partial Nodule Removal On Nitrogen Fixation And Regrowth Of Phasey Bean
- The Long Term Effect Of Lime On Soil PH
- Pastures
- The Management Of Bent Grass (agrostis Tenuis) Dominant Pastures In Victoria Using Glyphosate
- Effect Of Time Of Sowing On The Establishaent And Development Of Surface Sown Phalaris Aquatica
- Epic A New Variety Of Tall Fescue
- Early Growth Pattern And Sod Sowing Of Temperate Pasture Grasses
- Germination And Growth Of Senecio Madagascariensis Poir.(fireweed), A Toxic Plant Of Pastures In Coastal New South Wales
- The Role Of C.A.M. In Improving Rangeland Productivity
- Performance Of Seven Lucerne Cultivars Growing In Saline Soils
- Plant Density Decline Of Dryland Lucerne During Establishment
- Lucerne And Sainfoin Production On Two Alpine Soils In East Gippsland, Victoria
- Plant Environment Studies In Regional Evaluation Of Pasture Species
- Factors Affecting Seed Production In Jemalong Barrel Medic Based Pastures At Roseworthy Agricultural College
- The Need For Change In Making The Best Use Of Medics In The Cereal Livestock Farming Systems Of South Australia
- An Evaluation Of Mowing Trials
- Subterranean Clover Improvement In New South Wales
- The Growth Of Trifolium Montanum In South Eastern Australia
- Caucasian Clovers For High Country : Future Research Perspectives
- Legume Evaluation In The Monaro Region Of New South Wales
- A Comparison Between Leucaena Leucocephala Cv. Peru And Acacia Angustissima CPI 51651 In South East Queensland
- Fitzroy Stylo A Pasture Legume For Northern Brigalow Soils
- Desiccation Resistance In Macrotyloma Axillare
- Herbicide Banding To Aid Establishment Of Sod Sown Pastures In Southern Queensland
- Palatability And Cultivar Selection In The Eragrostis Curvula Complex
- Plant Breeding
- Plum Breeding In Queensland
- Some Basic Breeding Objectives For Dryland Chickpeas
- The Inheritance Of Seed Coat Impermeability Characters In Trifolium Subterraneum L.
- Breeding Lucerne For The Australian Environment
- Breeding Lucerne For Tolerance To Acid Soils
- Intra And Interspecific Crosses In Macrotyloma (papilionaceae)
- The Development Of A Scorch Resistant Subclover
- Recent Advances Cell Culture In Plant Improvement
- Winter Wheats Towards Clarifying Current Confusion
- Maize Cultivars For The Eighties
- Early Maize For Cool Summer Areas Of Southern Australia
- Evaluation Of Pearl Millet As A Potential Grain Crop For Eastern Australia
- Tolerance Of Barley Genotypes To Manganese Deficient Calcareous Soil On Eyre Peninsula
- Influence Of Plant Type On Grain Yield In Rice
- Selection For, And Characteristics Of, Wheat With Determinate Tillering
- Role Of Plant Breeding In Changing Cropping Practices
- Summer Crops
- Chilling And The Germination And Early Growth Of Sorghum And Cotton Seedlings
- Grain Yield Of A Hybrid Sorghum And Its Parents
- Plant Characteristics Associated With High Grain Yield Of Sorghum
- Uniformity Of Spacing, And Grain Yield, In Sorghum
- Row Spacing And Plant Population Effects On Water Soluble Carbohydrates Yield In Sweet Sorghum Cv. Rio
- The Effect Of Row Spacing On Stem Volume And Its Relationship To Water Soluble Carbohydrate Storage After Anthesis
- Peanut As A Dual Purpose Crop
- Dry Season Soybeans In Northern Australia
- Late Senescence Oil Losses In Soybeans
- Response Of Soybeans To Simulated Insect Attack
- Crop Evapotranspiration And Soil Water Depletion Of Dryland Sunflowers
- The Estimation Of Sorghum Yield Following Grain Losses Due To Midge Damage
- Contrasting Effects Of Population Density On Yield Of Irrigated Grain Sorghum In The Ord River Irrigation Area
- Comparative Yields Of Turnips, Sugar And Fodder Beet On The Southern Tablelands Of New South Wales
- Estimated Costs Of Producing Alcohol From Sugar Or Fodder Beet On The Southern Tablelands Of N.S.W.
- The Development And Use Of Solid Stream Nozzles For Aerial Application Of Molinate For Barnyard Grass Control In Rice Crops In New South Wales
- Developing New Crops
- Floret Sterility In Rice As Influenced By Low Temperatures
- Tillage
- No Tillage In Australia Some Implications From Overseas
- Infiltration And Soil Erodibility Benefits Of Conservation Tillage
- No Tillage Crop Production Research In Northern N.S.W.
- Why Cultivate?
- Practical Experiences With No Tillage Farming On Dryland And Irrigated Crop Production At Moree, NSW
- Better Lupin Husbandry In Southern New South Wales
- Tillage Systems Yield Interactions With Previous History
- Effect Of Direct Drilling Techniques On Seedling Environment And Growth Response
- The Performance Of Cereals, Grain Legumes And Rapeseed On Stubble Versus Fallow In The Wimmera
- Direct Drilling In Southern New South Wales: A Means Of Increasing Productivity
- A Comparison Of Seedbed Preparation Methods For Wheat Production
- The Effect Of Fallow Length And Cultivation Method On Crop Development And Yield
- Adaptation Of Wheat To Direct Drilling
- Deep Ripping Experiments For Wheat
- Tillage Systems For Continuous Cropping In South Australia
- No Till Wheat Feasibility In Southern Queensland
- Conservation Tillage In Australia
- Winter Crops
- Survey Observations On Lupin Seed Quality
- The Effect Of Seeding Time And Density On Lupin Growth And Yield
- Grain Legume Research At Lincoln College
- The Yield Potential Of New Oilseed Rape Cultivars In Tasmania
- Yield Compensation In Safflower Following Hand Pruning To Simulate Frost Damage
- A Self Propelled Windrower For Trial Plots
- Differential Varietal Tolerance Of Wheat Cultivars To Herbicides
- Lodging In Barley Can Plant Growth Regulators Prevent It?
- The Influence Of Spatial Arrangement On Wheat Yield
- Responses Of Wheat Varieties To Vernalization And Daylength
- Kernel Number In Wheat: Effect Of Light
- Yield, Harvest Index, Grain Nitrogen And Phosphorus
- Effect Of Varying Densities Of Great Brome (bromus Diandrus Roth.) On The Growth And Development Of Wheat (triticum Aestivum L.) And Great Brome
- Suppression Of Lolium Rigidum In Wheat By Cultural Means
- Measuring Severities Of Foliar Diseases In Wheat Crops
- Yield Losses In Wheat Associated With Different Levels Of Resistance To Speckled Leaf Blotch (mycosphaerella Graminicola)
- Realization Of The Potential Of Triticale
- Aluminium Tolerance In Triticales
- The Effect Of Stubble Residues On The Germination And Early Growth Of Wheat
- Stubble Retention And Nitrogen Supply On Dryland Wheat Yield
- Cereal Vs. Non Cereal Crops On The Southern Tablelands Of N.S.W.
- The Effect Of Erosion On Grain Yield At Wagga Wagga
- Yield Limitations Of Irrigated Wheat In The Lower Namoi Valley
- The Effects Of Seeding Rate And Nitrogen Fertilizer On The Growth And Production Of Wheat
- High Input Wheat Trials At Muresk, W.A. 1980 81
- The Development Of Chickpeas As A Viable Grain Legume Crop For The Victorian Wimmera
- Agricultural Systems
- Conference Information
- Invited Addresses
- Invited Reviews
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Problems Of Maintaining Pastures In The Cereal Livestock Areas Of Southern Australia
- Background Papers: Problems Facing Australian Agriculture
- 1985 3rd AAC
- Closing Session
- Concurrent Sessions
- Cereal Nutrition
- Phosphords Efficiency In Wheat Cultivars
- Tactical Application Of Nitrogen Fertilizer To Wheat
- Varietal Differences In Response To Nitrogen Fertilizer
- Nitrogen Fertilizer Availability To Wheat Under Field Conditions In Queensland And Western Australia Measured With 15n
- Effect Of Management Technique On Soil And 15N Fertilizer Use By Wheat
- The Effect Of De Lamination Of Wheat During Grain Filling
- Field Test For Nitrate Uptake By The Crop
- Sulfur Status Of Soils Of The Darling Downs, Queensland
- Effect Of Cultivar, Seed Source And Manganese Supply On Growth And Manganese Uptake Of Wheat (T. Aestivum).
- Interaction Of Manganese Deficiency And Cereal Cyst Nematodes On Barley
- Efficient Utilization Of Phosphorus By Wheat
- Cereal Production
- Windrowing Of Grain Crops
- Management Techniques To Improve Barley Yield And Malting Quality In S.A.
- Yield Maximization In Malting Barley
- Performance Of Winter And Spring Wheats In A Cool Region
- Variation In The Susceptibility Of Wheat To High Temperature
- The Optimum Crop Develoment Pattern For South West Victoria
- Predicting Cereal Crop Yields For Rainfall Analysis In Semi Arid Dryland Farming Areas
- Effect Of Lupin, Peas And Barley On A Subsequent Wheat Crop
- Early Sowing Of Oats For Grazing And Recovery For Hay On Grain Production
- Conservation Farming And Stubble Management
- Conservation Farming Methods For Western New South Wales
- Conservation Farming In Northern N.S.W. A Six Year Case Study
- Conservation Tillage In Southern New South Wales
- Soil Conservation Effectiveness Of Stubble In Different Tillage Practices
- Effect Of Rotations And Stubble Treatments On Soil Fertility And Crop Yields
- Stubble Retention In N.E. Victoria
- Water Use By Weeds Growing In Stubbles
- Weed Response To Crop Stubbles
- Tobacco Crop Residue Management At Mareeba
- Management A Constraint To The Early Adoption Of Conservation Cropping In The South Burnett?
- Crop Legumes
- Some Factors Influencing Yield Accumulation In Pigeonpea
- Developmental Attributes Of Fababeans For Northern New South Wales
- Narbon Bean ( Vicia Narbonensis) A Potential Grain Legume For The Victorian Mallee
- The Performance Of Lupins On The Alkaline Grey Clays Of The Wimmera Region Of Victoria
- Some Preliminary Results On The Response Of Field Grown Lupins, During A Period Of Soil Water Depletion
- Performance Of New Field Pea Plant Types
- Intercropping Of Cassava With Different Soybean Varieties
- Drought Adaptation Studies On Soybeans In The Murrumbidgee Valley
- An Ideotype For Chickpea (Cicer Arietinum L.) In A Dry Mediterranean Environment
- The Performance Of A Guar Triticale Rotation On Deep Siliceous Sands In South East Queensland
- Breeding For Weather Resistance In Mungbean
- Crop Production
- Plant Spacing Research In Onions And Its Commercial Significance
- Processing Bean Variety Evaluation
- Yield Of New Brassica Cultivars
- A Comparison Of Brassica Forage Crops With Oats
- Evaluation Of Chinoli Rapeseed (brassica Campestris)
- Sowing Time Effects On Yield, Growth And Development Of Rapeseed In Central Western N.S.W.
- Flowering Date And Yield Of Rapeseed
- Response Of Rapeseed To Early Windrowing
- A Sweet Potato Time Of Planting Trial In Papua New Guinea
- Determination Of Sowing Times For Sod Seeded Crops On The Central Tablelands Of New South Wales
- Maturity Type Of Maize For Cool Summers In Southern Australia
- Grain Drydown Effects In Sorghum Hybrids Treated Preharvest With Glyphosate
- A Comparison Of Single And Multiple Row Planting Systems For Irrigated Grain Sorghum Production
- Maize/cowpea Intercrop: The Effect Of Varying The Relative Time Of Sowing Of The Component Crops
- Evaluation Of Intercropping For Fodder Production
- Yields And Yield Components Of Sunflower Hybrids
- Root Development In Grain Sorghum Hybrids
- The Growth And Development Of Pearl Millet As Affected By Photoperiod
- Effect Of Reduced Seed Set And Seed Position On The Oil Content And Mineral Composition Of Sunflower Seeds
- Influence Of Tobacco Cropping Frequency On (1) Tobacco Crop Yield And Cured Leaf Quality
- Influence Of Tobacco Cropping Frequency On (2) Soil Acid Extractable Phosphorus Levels In North Queensland
- An Approach To The Management Of Sub Tropical Tree Fruits And Nuts Based On An Understanding Of The Penological Cycle
- Reasons For The Lack Of Grower Adoption Of Technology Arising From Research
- An Approach To The Planning Of Projects To Improve The Rate Of Grower Adoption
- The Potential For The Use Of Growth Regulators In Poppy Production
- Deep Ripping And Drainage
- The Effect Of Deep Tillage And Fallow On Yield Of Wheat On A Hard Setting Soil
- The Effect Of Deep Ripping On Water Use Of Wheat In A Drought
- The Response Of Pastures To Deep Tillage And Superphosphate In NE Victoria
- Grain Yield, Yield Components And Growth Of Wheat Following Soil Amendment With Lime And Deep Ripping
- The Effect Of Lime And Ripping On Soil Manganese And Manganese In Wheat
- Effect Of Gypsum And Deep Ripping Treatments On Lucerne Productivity Under Spray And Flood Irrigation On A Red Brown Earth
- Waterlogging, Soil Aeration And Field Crop Response
- Drainage For Crops And Forages
- Waterlogging Of Wheat In The Field I. A Method For The Control Of Waterlogging And Drainage In The Field
- Waterlogging Of Wheat In The Field II. Root Growth And Crop Yield
- Deep Tillage Research On The Southern Tablelands Of NSW
- Emergence/establishment
- Seed Applied Herbicides And Antidotes For The Establishment Of Lucerne And Phalaris
- Nutrient Seed Coating Of Phalaris And Lucerne
- Effect Of Terra Sorb On Seed Losses Due To Ants
- Effect Of Terra Sorb On Establishment Of Pastures
- Pasture Establishment On Non Arable Land Dominated By Poa Labillardieri
- A Phytotonic Effect Of Endophyte On Ryegrass Establishment
- Hypocotyl Length As A Determinant Of White Clover Seedling Emergence
- The Effect Of Temperature On The Length Of Wheat Coleoptiles
- A Method For Determining The Location Of Wheat Seeds After Sowing And Its Application To The Evaluation Of Seeding Equipment
- A Technique For Evaluating Root And Shoot Competition In The Field
- Pasture Establishment Identifying The Weak Link
- Irrigation
- Simulations Assessing The Effects Of Sowing Date, Maturity Type And Irrigation Frequency On Wheat Yields In The Murrumbidgee Irrigation Areas.
- The Effect Of Irrigation Method On The Response Of Maize To Mineral And Organic Fertilizers
- Effect Of Root Size On The Water Relations And Growth Of Two Sorghum Cultivars
- Response Of Sunflower To Strategies Of Irrigation
- Effects Of Irrigation Duration On Seed Yield And Economy Of Sunflowers
- Water Use By Sunflower And Sorghum
- The Effect Of Saline Irrigation Water On The Growth Of Lucerne In Northern Victoria
- Managing Irrigated Lucerne For High Protein
- Effects Of Frequent Irrigation And Previous Gypsum Application On Light Interception And Growth Of Lucerne
- Lucerne Production Under Different Frequencies Of Flood Irrigation
- A Comparison Of Trickle And Interrow Surface Flow Irrigation Of Cotton I. Cotton Plant Development
- Comparison Of Trickle And Inter Row Surface Flow Irrigation Of Cotton II. Yield And Water Use
- A Comparison Of Trickle And Surface Flow Irrigation Of Cotton III. Gin Turn Out And Lint Quality
- Improving Irrigation Efficiency In A Semi Arid Sub Tropical Environment A. Cotton Yields And Water Use Efficiency
- Improving Irrigation Efficiency In A Semi Arid Sub Tropical Environment B. Application Efficiency
- Improving Irrigation Efficiency In A Semi Arid Sub Tropical Environment C. Leaf Expansion And Dry Matter Partitioning
- Cotton Cultivar Response To Irrigation Management In A Semi Arid Subtropical Environment
- Irrigation Of Green Beans
- The Effect Of Seedbed Particle Size On Wetting Patterns And Seedling Emergence Under Trickle.
- Growing High Yielding Rice With Less Water
- Nutrients In Runoff From Irrigated Pastures
- Nutrients In Runoff From Irrigated Crops
- The Development And Evaluation Of A Computer Based Management System For Irrigated Wheat
- Measurement And Recording Of Agronomic Data
- Agricultural Activity Sensed By Satellites
- Managing Electronic Capture Of Data From Field Trials
- Water Use And Carbon Dioxide Exchange Of Lucerne In Relation To Leaf Conductance And Leaf Water Potential
- Field Gas Exchange As A Component Of Agronomic Research
- Case Studies Of Irrigation Scheduling And On Farm Water Management In Eastern Australia
- Foliage Temperature Measurement And Its Application To Agronomic Research
- Nutrition
- Fertilizer Requirements Of Irrigated Linseed At Emerald II. The Interaction Of N, P And Zn With Long Fallow.
- Fertilizer Requirements Of Irrigated Linseed At Emerald Iii Fertilizer Type And Placement In The Field
- Irrigated Summer Fodder Crops Nitrogen Responses
- Response Of Irrigated Grain Sorghum To Split Application On Nitrogen. I. Time To And Duration Of Head Exsertion And Anthesis
- Response Of Irrigated Grain Sorghum To Split Application Of Nitrogen II. Biomass And Grain Yield
- Nitrogen Fertilisation Of Sole And Intercropped Sorghum And Soybean
- Relative Importance Of Soil And Fertilizer Nitrogen To Rice
- Optimising Rice Growth Under Continuous Cropping
- Alternative Nitrogen Fertilization Strategies For Rice Soils
- Responses Of Russet Burbank Potatoes To Applied Nitrogen
- Sugar Cane Zinc Deficiency Following Liming In Far North Queensland
- Fertilizer Requirements Of Irrigated Linseed At Emerald I. Nitrogen And Phosphorus.
- Persistence And Production Of Pasture Grasses
- Effect Of Drought On Subsequent Pasture Growth
- Growth Patterns Of Perennial Ryegrass Varieties
- Tillering Pattern Of Perennial Ryegrass Subject To Two Spring Grazing Pressures
- Tillering Pattern Of Early Maturing Perennial Ryegrass, And Spring Growth
- Productivity Of Perennial Pasture Species Through An Irrigation Cycle
- Perennial Ryegrass Selection For Improved Productivity Under Irrigation
- Secale Montanum Evaluated In The Alpine Pastures Of Eastern Victoria
- Secale Montanum (s. Dalmaticum Vis) A New Pasture Species For The South Eastern Highlands
- Persistance Of Perennial Ryegrass Cultivars In Southern Tasmania
- Pasture Nutrition
- Selective Tissue Tests For Defining The Phosphorus Status Of Wheat And Annual Pasture Legumes
- The Place Of Fertiliser Test Strips In Planning Pasture Fertiliser Programs
- Comparative Effectiveness Of ESPARP And Superphosphate As Fertiliser For Perennial Pasture
- Seasonal Variation In Soil P Levels
- Effect Of Boron Application On Herbage And Seed Yields Of Subterranean Clover
- Factors Affecting Plant Uptake Of Selenium Following Selenium Topdressing
- Strategic Grazing Of Selenium Top Dressed Pasture As A Means Of Selenium Supplementation For Sheep
- Recent Developments In Sulphur Bentonite Fertilisers
- Stability In Cation Concentrations During The Development Of Two Temperate Pasture Species
- Evaluation Of Phosphorus Efficient Pasture Species
- Pests And Diseases
- Virus Diseases Of Trifolium Species In Temperate Australia
- Barley Yellow Dwarf Virus In Tasmanian Pastures
- Studies Of Paspalum Scorch Disease In Victoria
- Physiological Responses Of Wheat To Speckled Leaf Blotch
- Breeding Disease Resistance Into Safflower
- Sugar Cane Poor Root Syndrome In Far North Queensland
- Inheritance Of Resistance To Fusarium Wilt Race 3 In Tomato
- A Needle Nematode (paralongidorus Sp.) Causing Poor Growth Of Rice In North Queensland
- Implication Of Pratylenchus Thornei In Wheat Yield Decline In Northern New South Wales.
- Simulated Insect Damage To Soybean Growing Tips
- Simulated Insect Damage To Soybean Leaves
- Hessian Fly : A Potential Threat To Stubble Retention In Australia
- Soil Acidity And Liming
- A Study Of Factors Associated With Soil Acidity Under A Legume Based Pasture In East Gippsland
- Growth Of Subterranean Clover With Lime And Fertilizer Molybdenum
- Effect Of Lime On Mineral Nutrition Of Subterranean Clover On Acid Soils
- Inoculation And Nodulation Of Subterranean Clover On Acid Soils
- Enumeration And Distribution Of Rhizobium Trifolii In An Acid Soil And Implications For Nodulation Of Subterranean Clover.
- The Effect Of Lime On Growth And Phosphorus Uptake Of Red, White And Subterranean Clover On An Acidic Soil
- A Comparison Of Lime And Sewage Ash To Correct Soil Acidity
- Breeding Phalaris For Greater Tolerance To Acid Soils
- Response Of Onions To Lime
- Sunflower Response To Acid Soil Factors
- Soil Acidification In N.E. Victoria
- Soil Management
- The Effect Of Erosion On Wheat Yield On Two Soil Types.
- Fertility Decline In Queensland Cropping Soils And Its Implications For The Future.
- Opportunities To Reduce Dryland Salting In South Eastern Australia
- Effect Of Topsoiling, Fertilisers, And Soil Conditioners On Pasture Production From Areas Where Topsoil Is Removed During Land Forming
- Soil Management For Intensive Cropping On Krasnozems
- A Survey Of Soil Bulk Densities Occurring Under Grazed And Ungrazed Situations In NE Victoria
- Management Of Some Strongly Duplex Soils In SE Tasmania
- Crop Production On Duplex Soils After Landforming
- Crop/pasture Farming Systems To Conserve Soil Resources
- Sub Clover And Other Pasture Legumes
- An Extension To The Geographical Sources Of Hardseededness In Persian Clover
- Balansa Clover A New Pasture Species
- Variability In Six Major Characteristics Of 12 Annual Medic Species
- Medics For The Victorian Mallee And Wimmera Progress And Future Needs
- Genetic Control Of Margers In The Annual Medic Cv. Sephi
- The Effects Of Cereal Straw On Productivity Of Annual Medic Pastures
- Evaluation Of Yellow Serradella Accessions In Southern Victoria
- A Progress Report On Lucerne Breeding In South Australia
- The Effect Of Two Harvesting Schedules On The Yield Of Lucerne Varieties With A Range Of Winter Dormancy Ratings
- Lucerne Persistence Under Dryland Grazing And Haycutting
- Coumestrol Content Of Lucerne In The Central West And Hunter Valley Of New South Wales
- Annual Rates Of N2 Fixation By Pasture Legumes On The Central Plateau Of Tasmania
- The Potential Of Trifolium Subterraneum Subspecies Brachycalycinum On Red Brown Earth Soils
- Options For Improving Subterranean Clover Pastures In Short Term Leys
- A Survey Of Seed Reserves Of Subterranean Clover Pastures On Southern Tablelands Of New South Wales
- An Interim Report On The Performance Of A Binary Mixture Of Subterranean Clover Strains.
- On Farm Establishment Of Trikkala Sub Clover In South West Victoria
- The Poor Subterranean Clover Stands Of Dairy Pastures In The Adelaide Hills
- Effects Of Irrigation Management On The Productivity Of Subterranean Clover Pastures
- The Effects Of Irrigation Management On Seed Production And The Regeneration Of Subterranean Clover
- The Productivity Of Three Annual Clovers (T. Balansae, T. Resurinatum And T. Subterraneum) Under Irrigation In Northern Victoria
- Relationship Of Diet Quality To Green Herbage Available On Lucerne Subterranean Clover Pastures
- Liveweight Gain Of Lambs On Lucerne, Other Legumes And Grass Sub Clover Pasture Over Winter
- Genetic Diversity Within Persian Clover (trifolium Resupinatum)
- Conservation Tillage/cropping
- Reduced Tillage On The Darling Downs
- The Effect Of Controlling Sorghum With Glyphosate On Tillage Practices And Soil Water Status
- Direct Drilling And The Early Growth Of Wheat
- The Effect Of Conventional Cultivation And Direct Drilling On Soil Temperatures During The Early Growth Of Wheat
- Nitrogen Fixation, Growth And Yield Of Summer Crops Under Two Tillage Systems
- Low Cost Seeder Conversions For No Till Farming
- Direct Drilling Ryegrass Into Kikuyu Effects Of Machine, Sward Preparation And Timing
- Factors Affecting Silvergrass (Vulpia Spp) Control With Direct Drilling
- Opportunity Cropping : A Concept For More Efficient Agriculture In Northern New South Wales
- Weeds And Weed Control
- Effect Of Various Herbicides On Pasture Production And Weed Control In A Spray Graze System
- Effect Of Bent Grass (agrostic Tenuis) Control In Pastures On The Wool Production Of Wethers
- Grazing And Herbicide Manipulation Of Pasture Composition The Effects On Direct Drilled Cereal Production
- Accumulation Of Nitrate In Silybum Marianum
- Control Of Gorse (Ulex Europaeus L) With Triclopyr Both Alone And In Combination With Picloram.
- Control Of Gorse Regrowth By Angora Wethers
- Competitiveness Of Toad Rush In Wheat
- Assessment Of The Weed Potential Of Rubus Cultivars Of Horticultural Significance
- The Tolerance Of A Range Of Cucurbit Varieties To Tank Mixtures Of Prefar And Alanap
- The Virtues Of Some Weeds On A Midlands Grazing Property
- Cereal Nutrition
- Conference Information
- Invited Addresses
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- Systems Approaches For Crop And Pasture Research
- 1987 4th AAC
- Contributed Papers
- Computer Systems
- Aspects Of The Development And On Farm Evaluation Of Siragcrop, A Computer Based Crop Management System
- Expert Systems: Potential To Provide Weed Control Advice
- The Development Of A Prototype Expert System For Agricultural Extension
- Computer Aided Management Of Sheep Flocks In Western Victoria
- PADFERT A Computerised Aid To Whole Farm Fertiliser Management
- Development Of A Crop Information Service
- Performance Of Two Wheat Crop Models Compared With The Australian Wheat Field Trial Database.
- Crop Production
- Stem And Ear Growth In Wheat And Its Effect On Grain Yield
- Effect Of Sowing Date Upon Yield. Growth And Development Of Weak And Strongly Vernalizing Wheat Cultivars.
- Variation Among Spring Wheats In Winter Feed Production : An Apparent Association Between Growth And Floral Development
- Lime Application Increases The Risk Of Take Fall
- The Effect Of Water Stress On Yield And Dry Matter Production Of Rapeseed
- Variability In The Duration Of Flowering In The Faba Bean
- Normal And Reduced Branching Lupins Have Similar Residual Value
- The Effect Of Various Planting Dates On Development And Yield Of Two Cultivars Of Chickpea
- Vegetative And Reproductive Characteristics Of Five Genotypes Of Mungbeans Grown Under Two Water Regimes
- Progress With Faba Beans In South East South Australia
- Grain Legumes For Low Rainfall Areas
- Narbon Bean A Grain Legume With Potential
- Effects Of Plant Density On Pigeonpea (Cajanus Cajan (L.) Millsp.) In Sub Humid Environments
- Adaptation Of Short Season Pigeonpea In Multilocation Yield Trials
- Productivity Of Conventional And New Pea Phenotypes In Victoria
- Plant Population On The Dynamics Of Rooting In Sunflower
- A Comparison Of Dryland Maize, Grain Sorghum And Sunflower In The Central Highlands Of Queensland
- Adapting Brassica Juncea To Southern Australia
- Long Season Barley Varieties For High Rainfall Areas
- Self Defence Chemicals Of Barley
- Managing High Yield Potential Barley Crops
- The Use Of Early Maturity To Increase Barley Yields On Light Soils In Low Rainfall Areas
- Growth And Water Use Of Normal And Reduced Branching Lupins
- Stem Reserves And The Response Of Barley To Elevated Post Anthesis Temperature
- Barley As A Dual Purpose Crop In Tasmania
- The Effect Of Metolachlor Herbicide On Toadrush And Ten Pasture Species During Establishment
- Control Of Spiny Burrgrass By Consol Lovegrass
- Chemical Fallowing In The Victorian Wimmera
- Tolerance Of A Range Of Crop Species To Seed Coating With Monocalcium Phosphate
- An Instrumented Boll Buggy For Cotton Trials
- Yield Potential Of Semidwarf Wheat And The Effect Of Lodging
- Extension And Economics
- Use Of Neighbourhood Groups For Developing Technology
- Decision Support For The Adoption Of Improved Pasture
- Farm Records Data Base For Management And Education
- The Relative Profitability Of Long Fallows In The Eastern Wheatbelt Of Western Australia
- Farm Machinery On The Central Highlands
- The Production Line Approach (PLA)
- Irrigation And Salinity
- Strategies For Irrigating White Clover With Saline Water On The Heavy Soils Of Northern Victoria
- The Performance Of Rapeseed On Salt Affected Soils Of The Wimmera Region Of Victoria
- Salt Tolerance Of Subterranean Clover Cultivars
- Inheritance Of Chloride Exclusion In Grapevines
- Salt Accumulation By Some New Citrus Hybrids
- Response Of Avocado To Irrigation With Saline Water
- Irrigation Scheduling Of Commercial Navy Beans
- The Effect Of Irrigation Frequency On Lucerne And White Clover Production In The Murrumbidgee Valley
- Soil Water Storage And Extraction For Irrigated Maize On A Red Brown Earth
- Potential Productivity Of Irrigated Maize In Northern Victoria
- Estimated Potential Of Cotton Grown Under Trickle Irrigation
- Factors To Consider When Setting Refill Points For Irrigation Scheduling
- Software For Irrigation Scheduling Using The Neutron Probe
- Trickle Irrigation Of Processing Tomatoes On Red Brown Earth: Effects Of Line Placement And Irrigation Strategy On Crop Productivity
- Water Use And Capillary Contribution To Irrigated Wheat At Griffith, N.S.W.
- Pasture Response To Amelioration Of Exposed Subsoils I
- Pasture Response To Amelioration Of Exposed Subsoils II.
- Limits To Pasture Performance In An Area Where Extra Soil Was Added During The Land Forming Process
- Response Of Malting Barley Grown On A Red Brown Earth To Spring Irrigation And Nitrogen Fertiliser
- The Response Of Irrigated Navy Beans To Applied Nitrogen
- Nitrogen Fertilizer Balance Of Irrigated Wheat Grown On A Red Brown Earth
- Nitrogen Accumulation From Late Applications Of 15 N Labelled Fertilizer
- Yield Limitations To Irrigated Grain Sorghum At Emerald, Queensland.
- Relationships Between N Supply, Crop Growth And Yield Of Irrigated Wheat
- Irrigated Crop Rotations On Beds In The Murrumbidgee Valley 1. Wheat Growth And Yield
- Irrigated Crop Rotations On Beds In The Murrumbidgee Valley 2. Summer Crop Growth And Yields
- Effect Of Temperature On The Photosynthesis Of Irrigated Pastures
- Photosynthesis Of Pastures Ponded With Irrigation Water
- Effects Of Irrigation And Nitrogen On Wheat Production On A Red Brown Earth
- Effects Of Irrigation On CO2 Exchange Of Wheat
- The Apetalous Flower Character In Rapeseed And Its Interaction With Irrigation
- Annual Clover Production Using Saline Irrigation Water
- Pasture Production
- Differing Winter Production Of Subterranean Clover Cultivars
- Growth Responses Of Subterranean Clover To Temperature
- Sub Clovers For Tasmania: Production Of Early Mid Season And Late Groups
- Sub Clover For Tasmania: Flowering Times Of Registered Cultivars
- Persistence Of Mt. Barker Subterranean Clover
- Seed Yield Of Three Annual Pasture Legumes In Response To Water Deficits During Flowering And Seed Development
- Potential Of Trifolium Subterraneum Ssp. Brachycalycinum For Northern N.S.W.
- The Potential Of Medicago Murex As A New Legume Species For The Eastern Wheat Belt
- Impact Of Grazing By Sheep On Subterranean Clover Cultivars
- The Effect Of Grazing On The Seed Production Of A Range Of Annual Medic Species
- Effects Of Defoliation On The Productivity Of A Persian Clover Wimmera Ryegrass Sward
- Leaf Growth And Apical Development In Perennial Ryegrass In Winter
- Establishing Phalaris In The Upper Hunter By Aerial Seeding
- Tillering And Its Influence On Seed Yield In Kangaroo Valley Perennial Ryegrass
- Effect Of Season And Sowing Method On Pasture Establishment On The Northern Tablelands
- A Burn/graze Strategy For Improving Natural Pastures In Northern New South Wales
- Persistence And Production Of Native And Exotic Warm Season Perennial Grasses At Walgett, North West NSW
- Fertilizers For Animal Production
- Mineralization Of Pasture Residues
- Determining Pasture Fertilizer Requirements By Satellite
- Wool Production From Pasture Biological And Economic Limits Within Western Victoria
- Change In Soil Nitrogen Under Established Swards Of Caucasian Clover
- Seed Yield Of Annual Pasture Legume Rhizobium Symbioses
- The Effect Of Fungicide And The Control Of Leaf Disease In Lucerne
- Tolerance Of Persian Clover Seedlings To Broadleaf Herbicide
- Performance Of Grasslands Hamua X Moroccan Red Clover Crosses In A Marginal Environment
- The In Vitro Digestibility Of Dead Pasture Grasses And Their Morphological Components
- Potential Of Tagasaste As A Forage Plant In A High Rainfall Temperate Environment
- Competition As A Factor In Sub Clover Cultivar Replacement
- Comparative Establishment Levels Of Old And New Pasture Species In Southern NSW
- Effects Of Depth Of Sowing Medic Seeds On Emergence Of Seedlings
- Medic Seed Distribution In Soil Profiles
- Seedling Vigour And Seasonal Growth Of Two Hardseeded Lines Of Persian Clover In South West Victoria
- A New Inoculant For Persian Clover (Trifolium Resupinatum) Cv Maral
- Shrubby Stylo Grazing Development Trial "Carfax" Brigalow Area III – Queensland
- Emergence And Early Growth Of Lotononis Bainesii
- Improved Permanent Forages For Dry Season Grazing In Northern Australia 1. Ponded Pastures
- Improved Permanent Forages For Dry Season Grazing In Northern Australia 2. Oversowing Adapted Species Into Native Pastures
- Improved Permanent Forages For Dry Season Grazing In Northern Australia 3. Browse Shrubs And Fodder Trees
- Establishment Of Surface Sown Pastures On Cracking Clays Near Walgett
- Establishment And Development Of Surface Sown Pastures On Black Earths Near Coolah
- The Effect Of Autumn And Winter Grazing Pressure On End Of Spring Pasture Production
- Assessing The Hydrogen Cyanide Potential Of Forage Sorghum
- Subterranean Clover Seed Yields Of Pastures In The Hamilton District
- Introducing New Cultivars Of Subterranean Clover Into Existing Clover Pastures In South West Victoria By Direct Drilling
- Adoption Of Higher Sowing Rates Of Subterranean Clover In South West Victoria
- Sowing Pastures By Direct Drilling In South West Victoria
- Imbibition As A Factor Affecting Germination In Subterranean Clover (Trifolium Subterraneum)
- Plant Breeding And Evaluation
- Ideotypes For The Central Wheatbelt Of W. A. Part II Wheat
- Ideotypes For The Central Wheatbelt Of W. A. Part III Grain Legumes
- Selection For Delayed Leaf Senescence To Improve Drought Resistance In Sorghum
- The Response Of Uniculm And Tillered Barley To High Seeding Rates
- The Possible Origins Of Presumptive Somaclones In Barley Tissue Cultures
- Sironaria And Sirothora Disease Resistant Safflower Cultivars
- The Field Performance Of Three Aphid Resistant, Harbinger Backcross Lines
- The Breeding And Selection Of An Aphid Resistant Harbinger Type Strand Medic By Backcrossing
- Regrowth Of Subterranean Clover Lines Differing In Plant Morphology
- Identification Of. Couchgrass (cynodon Spp) Cultivars For The Turf Industry Using Gel Electrophoresis Of Leaf Perosidase Enzymes
- The Domestication Of Danthonia Linkii, A Native Australian Grass
- Ideotypes For The Central Wheatbelt Of W.A. Part I Approach
- Plant Nutrition And Fertiliser Use
- Changes In The Response Of Wheat To Nitrogen Fertilizer In Northern New South Wales
- The Effect Of Four Nitrogen Fertilizers On N Use Efficiency (NUE) Of A High Yielding Barley (var. Triumph)
- Effect Of N Management On Rice Grain Maturation
- N Topdressing Requirements For Rice
- Nitrogen Fertilizer Timing And Placement For Aerial Sown Rice
- Effect Of Seed Structure On The Tolerance During Emergence Of Five Grasses To Phosphorus Seed Coating.
- Comparison Of Soil With Leaf Analysis For Predicting The Response Of Pasture To Superphosphate
- The Annual Requirement Of Pasture For Phosphate
- Variation Among Sunflower Genotypes In The Delay In Flowering Caused By P Deficiency
- Effects Of Fallow Length On Sorghum And Sunflower Responses To P
- Variation In Responses Of Barleys To Applied Nitrogen
- Influence Of Nitrogen On The Yield, Water Use And Water Deficits Of Wheat Grown In A Mediterranean Climate
- Influence Of Post Seeding Application Of Nitrogen On Grain Yield And Water Stress Of Wheat In A Mediterranean Climate
- Effect Of External Nitrate Supply On Nodulation Of Lupins 1.Growth And Nodulation Of Lupinus Angustifolius And Lupinus Albus
- Effect Of External Nitrate Supply On Nodulation Of Lupins II. Comparison Of Eighteen Lines Of Lupinus Angustifolius
- Effects Of Relief Or Onset Of Nitrogen Stress On Yield Components Of Linseed
- Liming, Water Deficits And C2h2 Reduction By Subterranean Clover
- Soil Acidification Under Permanent Pastures In NE Victoria
- The Effect Of Lime On Establishment, Nodulation And Herbage Yield Of Persian Clover In South West Victoria
- Presence Of Plant Secreted Flavones And Expression Of Nodulation Genes Of Rhizobium Trifolii In Acidic Conditions.
- Yield And Seasonal Production Of A Subterranean Clover Based Pasture Growing In An Acid Soil
- Foliar Micronutrients And Lime On Wheat Production
- Copper Deficiency Of Meat In Victoria. I. Cause
- Copper Deficiency Of Wheat In Victoria. II. Correction
- Effects Of Applying Sulfur To Sulfur Deficient Linseed And Rapeseed
- Response Of Wheat To Foliar Applied Copper On Five Hamilton Soil Types
- Soil And Plant Analysis For The Prediction Of Copper Deficiency In Wheat.
- Variability Of Rice Yield Responses To Nutrients On Farms In The Central Philippines
- Boron Tolerance Of South Australian Wheat Cultivars
- Effect Of Recent Cropping History On The Nitrogen And Phosphorus Requirement Of Maize
- Phosphorus Uptake By Phalaris From 32p Labelled Seed Coatings And Drilled Granules
- A Fertiliser Model For Predicting Potato Yields Based On Soil Tests
- Potassium Requirements Of Potatoes Grown On Krasnozems In Tasmania
- Cereal Whole Tops Plant’ Analysis Methods Used By CSBP
- Effect Of Superphosphate On Relative Production From Trifolium Subterraneum And T. Balansae Pasture At Cranbrook, Western Australia
- Concentration Of Copper In Subterranean Clover Compared To Polymorpha Medic Under Field Conditions
- Deep Placement Of Ammonium Type Nitrogen Fertiliser In South West Victoria
- Soil Tillage And Soil Water
- Evaluating Stubble Management Techniques
- Preseason Management For Weed Control In Direct Drilled Crops
- Stubble Retention Effects On Soil Water Storage And Yield
- Effects Of Cultivation And Stubble Retention On Soil And Stubble Borne Pathogens Of Wheat In Victoria
- Broadcasting Versus Drilling Of Wheat At Five Seed Rates
- Soil Water And Thermal Infrared Determination Of Water Stress Of Wheat Growing On A Red Brown Earth In Southern New South Wales
- The Infrared Thermometer As A Practical Irrigation Scheduling And Crop Management Tool
- Co2 Balance Of Sunflower Subjected To Moisture Stress During Grain Filling
- Physiological Variation Contributing To Genotype By Environment (water Supply) Interaction In Barley
- Osmotic Adjustment In Three Pasture And One Grain Legume In Response To Severe Water Deficits During Flowering
- Adaptation Of Upland And Lowland Rice To Soil Moisture Deficits
- Relationships Between Water Use, Yield And Grain Size Of Barley I. Yield
- Relationships Between Water Use, Yield And Grain Size Of Barley II. Grain Size
- Residual Water In Sandy Soils After Various Rotations And Agronomic Treatments
- Effects Of Wheat Residues And Tillage On The Water Balance Of A Red Earth Soil
- Variation In Water Use And Dry Matter Production Among Peanut Cultivars Grown Under Moisture Deficit
- Water Use Of Wheat In The Victorian Mallee
- Amelioration Of Hardsetting Red Duplex Wheat Soils With Gypsum
- The Effect Of Soil Moisture Ameliorants On Pasture Establishment On Silicious Sands
- Stubble And Tillage Effects On Soil Characteristics And Grain Sorghum Yield
- Strategies And Tactics With Short Fallowing
- Reduced Tillage Vegetable Production Systems For Coastal New South Wales
- Extension Of Conservation Tillage Technology Onto The Farm South East Darling Downs, Queensland
- Precision Sprinkler For Small Field Plots
- Computer Systems
- Donald Oration
- Conference Information
- Invited Addresses
- Salinity An Environmental Constraint On Crop Productivity
- "The Shock Of The New" Constraints On Farmer Use Of New Technology
- Economic And Marketing Influences On Future Agronomic Production
- Soil And Plant Testing In Australia1
- The Genetic Organization Of Nitrogen Fixing Rhizobium Species: Past And Future Applications To Agriculture.
- Grain Legumes In Australia
- Water Use Efficiency Of Non Irrigated Field Crops
- Effects Of Deep Ripping On Cropping Soils And Crop Production
- Opportunities For Improving Crop Yields Through Research A Physiological Perspective
- Public Lecture
- Invited Reviews
- The Genetic Organization Of Nitrogen Fixing Rhizobium Species: Past And Future Applications To Agriculture.
- Grain Legumes In Australia
- Water Use Efficiency Of Non Irrigated Field Crops
- New Directions For Irrigated Pastures
- Effects Of Deep Ripping On Cropping Soils And Crop Production
- Soil And Plant Testing In Australia
- Workshop
- Contributed Papers
- 1989 5th AAC
- Contributed Papers
- Crop Production
- Light Interception And Growth By Wheat And Barley In Western Australia
- Variability In Wheat Growth On A Duplex Soil In Western Australia
- Tolerance Of Two Wheat Cultivars To 2,4 D In Relation To Their Ear Development Stage
- The Date Of Anthesis Is Not Important For South Australian Wheat Crops
- Wheat Yields On Acidic Wodgil Soils
- Water Stress And Wheat Crop Growth
- Two Tall Wheats Equal Or Outyield Two Semi Dwarfs With Late Sowing
- Flowering Times Of Wheat In Western Australia : A Model Approach
- Increasing Inter Row Spacing Decreases Wheat Yield
- Planting Density Effects On The Growth, Yield And Water Relations Of Wheat
- Silvergrass Residue Effects On Wheat
- Reduced Germination And Survival Of Wheat Seed Exposed To High (35 40ðc) Temperatures
- Soil Evaporation, Transpiration And Transpiration Efficiency Estimates For Old And Modern Varieties Of Wheat
- The Response Of Wheat To High Temperature During Grain Growth
- Frost Risk And Site Variation Therein
- A Comparison Of Six Methods For Estimating Crop Evapotranspiration
- Use Of A Sensitivity Analysis To Critically Evaluate A Model Of Cereal Growth
- Simulating The Western Australian Wheat Sheep System
- Phytotoxic Activity Of Barley
- Cereal Aphids In High Yield Cereal Crops
- Studies On The Molecular Forms Of Alpha Amylase And Their Implications For Improving Malting Quality In Barley
- Response Of Barley To Hydrogen Ion Toxicity
- The Effect Of Grazing On Grain Yield Of Winter Barley In Tasmania
- Oats As A Multi Purpose Crop In High Rainfall Environments
- Rye As An Alternative Cereal Under High Rainfall Conditions
- Grazing And Recovery Of Cereals In South Western Australia
- Restricted Tillering In Triticale Cv. Currency An Impediment To Grain Yield?
- Production Of Oats, Vetch And Medic Sown Alone And In Various Combinations
- Effect Of Time To Flowering On Yield Of Rapeseed For Drier Areas Of The South Western Australian Cereal Belt
- Lupin Yield Response To Plant Density In The Western Australian Wheatbelt
- Effect Of Solar Radiation And Temperature On Potential Maize Yield
- Rooting Depth Of A Range Of Crop Species On Sandhill Soils In The Victorian Mallee
- Development Of Rough Seeded Lupins For Agriculture
- Effect Of Seed Size On Lupin Establishment And Yield
- Contribution Of Flower And Pod Abscission To Reproductive Abortion In Lupinus Angustifolius
- The Sequence Of Flower And Pod Abscission On The Main Stem Inflorescence Of Lupinus Angustifolius
- Agronomic Management Of Field Peas On Acid Soils.
- Seeding Date And Rate Of Conventional And Semi Leafless Field Peas
- Factors Associated With Reduced Pea Yields In South Eastern South Australia
- Trifluralin Without Incorporation
- Evaluating The Concept Of Thermal Time
- A Mechanical Soil Sampling System To Encourage Increased Use Of Bioassay Tests For Soil Borne Cereal Diseases
- Removal Of Nutrients In Harvested Grain
- Grass Pasture Leys Benefit Peanut Production In Southern Queensland
- Effect Of Shading On Growth Of Two Peanut Cultivars
- Ceres Sorghum (sat): A Model Of Sorghum Growth And Development In The Semi Arid Tropics
- Time Of Sowing Of Irrigated Grain Sorghum In The Murrumbidgee Valley
- Sink Sink And Source Sink Interactions Between Stems And Reproductive Structures Of Sunflowers With Ontogeny And Water Stress
- A Model To Predict Growth And Yield Of Sunflower
- The Influence Of Sowing Date On Oil Quality Of Irrigated Sunflowers
- Cotton Yield Predictions For Lombok And Flores Indonesia
- Evaluation Of Sowing Opportunities For Lombok And Flores Indonesia
- Excess Growing Season Rain For Lombok And Flores Indonesia
- Soybean Irrigation Scheduling Trials In The South Burnett
- Limitations To The Yield Of Soybeans In A Tropical Environment
- Seed Treatments Improve Germination And Growth Of Soybean Cv. Chaffey Sown Into Cool Soils
- The Effect Of Paclobutrazol On The Growth Of Fiord Faba Beans
- Yield Responses Of Faba Bean To Inoculation
- Effect Of Varying Crop Population On Lentil And Weed Growth
- Effect Of Bees On Seed Yield And Components In Pigeon Pea
- The Effect Of Sowing Rate And Sowing Date On Yield And Yield Components Of Plantago Ovata (forsk.) In Northern Australia
- Floral Initiation Of Sesame As Affected By Photoperiod And Temperature
- Comparative Productivity Of Oilseeds And Cereals
- Development Of Indian Mustard As An Oilseed Crop
- Preharvest Factors Affect The Quality Of Broccoli After Storage
- The Simulation Of Flowering (bunch Emergence) In Plant Crops Of Banana (Musa Sp)
- The Flowering Of Carambola (averrhoa Carambola L) Is More Strongly Influenced By Water Stress Than By Photoperiod And Diurnal Temperature Variation
- Sugar Pulsing Of Cut Geraldton Wax And Kangaroo Paws
- Retaining Suppleness Of Dried Flowers And Foiliage With Humectants
- Cold Storage Of Geraldton Wax, Kangaroo Paw And Banksia
- The Relative Importance Of Seminal And Nodal Roots In Supplying Water And Nitrogen To Wheat
- Extension And Economics
- Using A Model To Assess The Profitability Of Applying Nitrogen To Dairy Pastures
- An Expert System For Extension Of Weed Control Advice
- Evaluation Of Written Extension Texts
- The Development And Evaluation Of Siragcrop Software For Irrigated Crops
- Padfert Extension Experiences
- Market Limitations To The Adoption Of New Crops A Case History From North West Europe
- Use Of Weather Based Spray Decision Models For Reducing Fungicide Application Costs On Peanuts In North Western Australia
- Irrigation, Drainage, Salinity And Waterlogging
- Effects Of Grazing Management On Productivity Of Pure Legume Pastures Grown Under Irrigation In Northern Victoria
- Effects Of Irrigation Frequency On The Productivity Of White Clover, Red Clover And Lucerne In Northern Victoria
- Cotton Irrigation Scheduling In The South Burnett
- The Effect Of Surface Soil Stability On Water Intake And The Yield Of Irrigated Maize
- Soybean Response To Moderately Saline Water
- Mineral Content In Seed Of Soybeans Grown Under Saline Conditions
- Comparison Of Three Techniques For Revegetating Salt Affected Land With Halophytes
- The Role Of Seed Placement And Gypsum In Improving The Establishment Of Spiny Bluebush On Scalded Duplex Soils In North Western Australia
- Subsurface Drainage Reduces Accessions To Watertables
- Moisture Relations, Groundwater Recharge And Dryland Salinity
- Adverse Effects Of Waterlogging On Germination And Survival Of Wheat, Barley And Rice Seed
- Effect Of Waterlogging On Photosynthesis And Stomatal Conductance Of Chickpea Leaves
- Variation In Water Use Efficiency And Its Correlation With Carbon Isotope Discrimination In Peanut Cultivars Grown In The Field
- Using Carbon Isotope Discrimination To Predict Water Use Efficiency
- Waterlogging Affects Tme Water Relations Of Wheat Leaves
- Effect Of Waterlogging On The Growth And Yield Of Sweet Potato (Ipomoea Batatas L.)
- Changes In Concentrations Of Oxygen, Carbon Dioxide And Ethylene In Soil Solutions During Waterlogging Of Some West Australian Soils
- Adverse Effects Of Waterlogging On Growth Of Lupin And Field Pea Cultivars
- Water Use Of Plum Trees (prunus Salicina) Trained To Four Canopy Arrangements
- Salinity A Possible Tool For Improving Tomato Quality?
- A Possible Physiological Mechanism Underlying The Response Of Sugar Accumulation In Rockmelons Ripened Under Reduced Irrigation
- Growth Responses Of Annual Legumes On Saline Soils
- Pasture Production
- Effects Of Moisture During Flowering On Seed Yield Of Subterranean Clover
- Competition Between Mt Barker And Karridale Sub Clover. Sown In Binary Mixtures
- Establishment Of Subterranean Clover Under Direct Drilled Cereal Crops
- Losses Of Subterranean Clover Seed From Dry Pasture Residues During Grazing By Sheep In Summer Autumn
- The Sub Clover Status Of Permanent Pastures In The Lower Rainfall Areas Of Tasmania
- The Fate Of New Sub Clover Cultivars In Old Mt. Barker Pastures In Tasmania
- Spring Sown Sub Clover: Will It Regenerate?
- Tolerance Of Sub Clover Cultivars And T. Balansae Cv Paradana To A Range Of Herbicides In Tasmania
- Hard Seed Reserves Of Subterranean Clover At Various Soil Phosphate Levels
- Comparative Production Data From Commercial And Non Commercial Subterranean Clover Lines
- The Relationship Between Sowing Rate, Seedling Density. And Seed Yield Of Trikkala Sub Clover On Farm Paddocks In South West Victoria
- Effects Of Defoliation On Subterranean Clover Seed Production
- A Comparison Of The Root Structure Of Trifolium Repens Cvv. Huia And Haifa
- Root Shoot Interactions In Irrigated White Clover
- Perennial Pasture Legumes In The Murrumbidgee Valley
- Effects Of Temperature On The Germination Of Trifolium Resupinatum And T. Balansae
- Competitiveness Of Annual Forage Clovers With Capeweed
- Preliminary Evaluation Of Persian Clover
- The Role Of Balansa Clover (trifolium Balansae) In Cool Elevated Pastures
- Comparative Production Of Annual Medics In Jordan And South Australia
- Growth Responses Of Annual Medics To Temperature
- Longevity Of High Temperature Embryo Dormancy In Medicago Murex And Trifolium Subterraneum
- Medic Seed Dynamics Through Crop/Pasture Rotations
- Seed Yield From Medicago Murex Grown Alone And With Subclover
- Saprophytic Competence In Four Acid Soils Of Strains Of Rhizobium Meliloti
- Santiago An Improved Variety Of Burr Medic
- Importance Of Sampling Time In Assessing Germinability Of Burr Medic
- Water Use By Serena Medic (medicago Polymorpha) On Contrasting Soil Types In The Eastern Wheatbelt Of Western Australia
- Responses Of Cultivars Of Yellow Serpadella (ornithopus Compressus) To Inoculation With Strains Of Rhizobium Lupini
- Pasture Legume Seed Survival Following Ingestion By Sheep
- Predicting The Emergence Of Annual Pasture Legumes: Commercialisation Of A Soil Coring Technique
- Annual Pasture Legume Growth On Acid Soils I The Performance Of Yellow Serradella (Ornithopus Compressus Cv. Toro)
- Annual Pasture Legume Growth On Acid Soils II The Root Growth Of Seedling Subterranean Clover (Trifolium Subterraneum Cv Junee) And Barrel Medic (Medicago Truncatula Cv. Paraggio) On An Acid Soil Following Deep Tillage And Deep Lime Placement
- Annual Pasture Legume Growth On Acid Soils Iii Effect Of Soil Acidity On The Growth Of Seedling Subterranean Clover (trifolium Subterraneum Cv. Junee) And Barrel Medic (medicago Truncatula Cv. Paraggio)
- Loss Of Surface Sown Perennial Grasses In Their Establishment Year In North Western New South Wales
- Performance Of Perennial Ryegrass In Marginal Environments In South Western Victoria
- Effect Of Seed Structure On The Water Relations Of Germinating Pasture Grass Seeds
- Phalaris Establishment Sowing Alternate Rows To Overcome Perennial Ryegrass Competition
- Root Distribution Of Consol Lovegrass
- Possible Alleopathy In Consol Lovegrass
- Alternative Perennial Grasses For Acid, Low Phosphorus Fertility Soils In North Eastern Victoria
- Seed Production Of Curly Mitchell Grass (astrebla Lappacea)
- Effects Of Paclobutrazol Treatment On The Growth And Seed Yield’ Of Phalaris
- Establishment And Tillering Of Phalaris And Cocksfoot On An Acid Soil
- Effect Of Broad Leaf Herbicides On Production Of Annual Legumes
- Pasture Seedlings Respond To Fungicides On Northern Tablelands Of N.S.W.
- Emergence And Survival Of Sown Perennial Grasses In A Semi Arid Environment
- Establishment Of Surface Sown Perennial Pastures To Replace Annual Grass Weeds
- Pre Season Management Of Pasture And Its Influence On Annual Grass Regeneration
- The Status Of Pasture Legumes In Central NSW
- Effects Of Methods Of Sowing On Establishment Of Forage And Pasture Species
- The Structure Of Managed Tagasaste (Chaemecytisus Palmensis)
- Potential Intake Of Tagasaste (chaemecytisus Palmensis) As Influenced By Shrub Structure
- Medic Pasture Composition Changes Over Three Years Of Chemical And Physical Manipulation At Merredin, Western Australia
- Plant Breeding And Evaluation
- Ranking Sub Clover Lines With Respect To Several Characteristics Considered Simultaneously
- Breeding Yanninicum Subclovers W1th Improved Clover Scorch Resistance
- Evaluating Selection Criteria To Improve Salt Tolerance In White Clover (trifolium Repens L.)
- Rosedale A New Brachycalycinum Subclover
- Native Grasses On The Southern And Central Tablelands Of N.S.W. Potential For Utilization
- Persistence Of Temperate Perennial Grasses, Northwest Slopes, NSW
- Responses Of Perennial Lotus And Astragalus Accessions To Acid Soils
- Stem Nematode Effects On Lucerne Cultivars
- Identification And Characterisation Of Water Limiting Environments Within Queensland Wheat Breeding Trials
- An Investigation Of The Adaptation Of Seleci Ed Cimmyt Wheat Germplasm To Water Limiting Environments In Queensland
- Limitations Of The Use Of Drought Susceptibility Indices Based On Relativeyield For Selection Of Drought Adapted Genotypes
- A Classification Based Approach For Characterising Genotype Drought Adaptation For Selection For Stress Adaptation
- Vernalisation Requirements Of Contrasting Wheat Genotypes
- Grain Mass Variation In Wheat
- Contribution Of Genetic Change To Yield Improvement In Australian Spring Wheats
- Morphological And Physiological Characters Associated With Higher Grain Yields Of Modern Wheats
- Root:shoot Ratios In Old And Modern Australian Wheats
- Increasing Wheat Yields On The South Coast Of Western Australia
- Selection Techniques For Improved Yields And Drought Resistance Of Wheat
- Agronomic Characteristics Of Five Spring Wheats
- Genotypic Variation In The Response Of Barley Cultivars To Different Levels Of Phosphorus Fertility
- An Evaluation Of Lines Of Uniculm Barley
- A Rapeseed Ideotype For High Yields
- Thermal Requirements Of Three Brassica Species Under Various Seeding Dates
- Yield By Environment Interactions For Warm Season Perennial Grasses
- Variation In Soil Water Extraction And Harvest Index Among Peanut Cultivars In Response To Drought
- Genotype Environment Interactions And Yield Stability In Rice Grown Under Dryland Conditions
- Yield Potential Of Short Duration Rice Varieties In South Western New South Wales
- Differential Performance Of Diverse Sorghum Hybrids Over A Wide Range Of Environments
- Genotype By Environment Interaction For Radiation Use Efficiency In Grain Sorghum
- Genotype By Environment Interaction For Transpirat1on Efficiency In Grain Sorghum
- Variation In Carbon Isotope Discrimination Among Diverse Sorghum Hybrids
- Early Maturing Navy Beans For Tasmania
- A Screening Technique For Weathering Resistance In Mungbean
- Development Of An Ideotype For Weathering Resistance In Mungbean
- Functional Ideotypes For Increasing And Stabilizing Chickpea Yield
- An Initial Evaluation Of Fifty Diverse Lines Of Lentils (lens Culinaris) For Adaptation To South East Queensland
- Introduction And Evaluation Of Pasture Legumes In High Rainfall North Western Tasmania
- Plant Nutrition And Fertiliser Use
- Variation In Phosphorus Content Between Subspecies Of Subterranean Clover
- Remote Sensing And Tme Phosphorus Status Of Pasture
- Agronomic Requirements Of A New Semi Dwarf Rye Variety 1. Phosphorus And Sowing Rate
- Agronomic Requirements Of A New Semi Dwarf Rye Variety 2. Nitrogen And Phosphorus
- Subterranean Clover Decline And Phosphorus Nutrition
- Residual Value Of Phosphate For Lupins Increased By Deep Placement
- Comparative Phosphate Requirements Of Burr Medic, Yellow Serradella And Subterranean Clover
- An Assessment Of The P0tential For Direct Application Reactive Phosphate Rock In Australasian Agriculture
- Phosphorus Requirements Of Wheats And Barleys Grown In South Australia
- Effect Of Nitrogen And Phosphorus Seed Coating On The Emergence Of Tall Fescue
- Early Growth Response Of Tall Fescue To Nitrogen And Phosphorus Seed Coatings
- Residual Nitrogen And Wheat Production
- Rapid Sap Test For Monitoring The Nitrogen Status Of Crops
- Determining The Nitrogen Requirement Of Crops: 1 Wheat
- Determining The Nitrogen Requirement Of Crops: 2 Oilseed Rape
- Determining The Nitrogen Requirements Of Crops : 3 Sunflowers
- Predicting The Yield Response Of Wheat To Topdressed Nitrogen
- Tests To Predict Yield Response Of Wheat To Topdressed Nitrogen
- The Effect Of Nitrogen Fertilizer And Plant Population On Growth And Yield Of Minicorn
- Leaching And The Plant Availability Of Organic Nitrogen
- Applying Nitrogen To Late Sown Wheat Is A Waste Of Money In South Australia
- Effect Of Split Nitrogen Application On Yield And Protein Content Of Wheat In Northern New South Wales
- Impact Of Payment For Quality On Nitrogen Recommendations For Wheat
- Long Term Uptake Of Fertilizer Nitrogen Residues By Wheat
- Fate Of 15n Urea Applied To A Wheat Crop As Affected By Stubble Management
- A High Incidence Of Crown Rot (fusarium Spp.) In Wheat Grown Under High Levels Of Nitrogen
- The Effect Of Nitrogen Fertilizer And Disease Control On Yield And Protein Content Of Wheat In Canterbury, New Zealand
- Effect Of Nitrogen, Gibberellic Acid And Population On Yield And Seed Quality Of Navy Beans
- Applied Nitrogen And Water Use Efficiency Of Rapeseed
- Nitrogen Application And Waterlogging In Wheat
- Zinc Affects Yield, Protein And Quality Of Wheat
- Boron Tolerance Of Australian Wheat Varieties
- Low Boron In Seed Depresses Soybean Seed Yield
- Grain Set Failure And Boron Deficiency In Wheat In Thailand
- Supra Optimal Manganese Suppresses The Effect Of Gaeumannomyces Graminis Var. Tritici On Grain Yield Of Wheat
- Seasonal Variation Of Aluminium, Manganese And Ph In Acid Soils
- Assessment Of Soil Tests For Aluminium Toxicity In Sub. Clover
- Ash Alkalinity Of Farm Produce
- Effects Of Liming On Legume Content Of A Dairy Pasture On The North Coast Of New South Wales
- Liming Reduces The Soil Strength Of A Krasnozem
- Effect Of Soil Texture On Injury To Crop Species By Seed Coating With Monocalcium Phosphate
- Soil, Tillage And Water
- Soil Strength Development In Loamy Sand Soils: Effects Of Compactive Forces And Multiple Wetting And Drying Cycles
- The Influence Of Stubble Management And Nitrogen Fertiliser On Nitrogen Fixation In Chickpea
- The Effect Of Wheat Stubble On The Emergence Of Burr Medic
- Management Options For Intensive Cropping Of Red Duplex Soils
- Effects Of Tillage On Burial Of Pasture Legume Seed And Seedling Emergence
- Tillage And Stubble Management Effects On Lupin Yields In A Long Term Wheat/lupin Rotation On A Red Earth Soil At Wagga Wagga
- Long Term Effects Of Rotation, Tillage And Stubble Management On Wheat Yields On A Red Earth Soil In Southern N.S.W.
- Fallowing For Moisture Conservation In The Victorian Mallee
- The Effect Of Deep Ripping And Gypsum Application On Sodic Duplex Soils For Pasture Production
- Deep Ripping Responses In Annual Pasture Legumes
- Reduced Cult1vation For Irrigated Maize On A Red Brown Earth
- The Effects Of Straw Mulch On Establishment Of Annual Medics
- Shallow Autumn Cultivation Improves Burr Medic Pastures
- Adoption Of Direct Drilling For Sowing Pastures In South–West Victoria
- A Versatile Seeder For Sowing Conventionally Tilled Or Direct Drilled Experimental Plots
- Tillage And Sowing Depth Affect Pleiochaeta Root Rot And Lupin Establishment
- Deep Ploughing Improves Nutrient Distribution And Crop Performance
- The Effect Of Crop Tillage On Pasture Regeneration And Production On A Hard Setting Red Brown Earth
- The Extent Of Acid Soils In The Cropping Region Of North Eastern Victoria
- A Stochastic Method For Simulation Of Leaching In Soil
- An Inexpensive High Resolution Weighing Lysimeter
- A Large And Low Cost Rain Out Shelter Design
- Yield Of Wheat After Direct Drilling With A Modified Combine
- Crop Production
- Donald Oration
- Conference Information
- Invited Papers
- Busselton Symposium
- Katanning Symposium
- Waterlogging: A Hidden Constraint To Crop And Pasture Production In Southern Regions Of Australia
- Relationships Between Cropping And Livestock On Mixed Farms In Southern Australia.
- New Discoveries In Pasture Research: Current Status & Developments With Temperate Perennial Pastures And Shrubs In Southern Australia
- Pasture Production From Saltland
- Roles For New Pasture Legume Species In Southern Australia
- Management Of Annual Legume Pastures
- Nutritive Value Of Pasture Species
- Coping With Waterlogging In The High Rainfall Zones Of Southern Australia
- Root Diseases As A Major Constraint In High Rainfall Cropping Systems
- Environment Of The High Rainfall Zone Of Southern Australia And Implications For Agriculture
- Merredin Symposium
- Roles Of Grain Legumes In Sustainable Dryland Cropping Systems
- The Development Of Pasture Legumes For The Low Rainfall Cereal Livestock Zone Of Southern Australia
- Dryland Soil Salinity Cure, Containment Or Catastrophe?
- Adapting Farming Systems Research Concepts To Australian Research Needs
- MIDAS: An Economic Modelling Approach To Determining Research Directions For Wholefarm Systems
- Development Of A Minimum Tillage System: Rutherglen Experience
- Developing A Research And Extension Philosophy For An Agricultural Region
- Competition Versus Co Operation In Crop Production Demand Enhancement Versus Supply Control
- Solutions To Problems Of Oversupply In Crop Agriculture
- Farming Systems Of Southern Australia
- Plenary Sessions
- Implications Of The Greenhouse Effect For Australian Agriculture
- Looking For Products, Looking For Markets
- Research And Extension
- Research And Extension: Delivering The Goods
- Using Modern Communications Technology: Turning On, Not Turning Off
- Future Funding For Research And Extension: Industry Research Council Issues
- Future Funding For Research And Extension: A State Department Of Agriculture Viewpoint
- Commercialization: The New Zealand Experience
- Contributed Papers
- ASA 1992
- Concurrent Sessions
- Alternative Practices And Plant Protection
- Euphorbia Esula: A Challenge For Non Herbicidal Weed Control In North America
- Potential Harmful Effects Of High Endophyte Content In Perennial Ryegrass On Companion Legumes
- Crop Rotation For The Control Of Wild Oats In Wheat
- Organic Rice Cultivation For Sustaining Irrigation Farming In The Riverina
- The Distribution Of Tannins In Lotus Spp.: Variations Within Plant And Stage Of Maturity
- Gramine: The Occurrence Of A Self Defence Chemical In Barley, Hordeum Vulgare L.
- Cropping Systems
- Effects Of Different Preceding Crops On The Growth, Yield Components And Grain Yield Of Winter Wheat And Winter Barley
- A 20 Year Comparison Of Four Agricultural Systems For Their Sustainability In The Northern Wheat Belt
- Long Term Rotation Studies The Influence Of Clover And Crop Sequence On Maize Yields At Glen Innes
- Chickpea Cultivar By Planting Time Studies In Queensland
- Maize Silage: Looking Back And Planning Ahead
- Soil And Water Management For Irrigated Crops On A Red Brown Earth Soil
- Irrigated Faba Beans From Two To Two Hundred
- An Irrigated Rice Based Cropping System For Tropical Australia
- Comparative Water Use Of Dryland Crop And Pasture Species
- An Evaluation Of Cereal Zone Rotations For Achieving Short Term Aims And Long Term Sustainability
- Grazing Resources Ecology
- Seed And Herbage Production And Cyanogenic Potential Of White Clover Cultivars Under Irrigation At Neuarpurr, Victoria
- A New Approach To Improving Yield Of White Clover Role Of Roots
- Subterranean Clover Persistence In South Western Victoria
- Population Dynamics Of Trifolium Subterraneum,, Medicago Murex And Trifolium Balansae Grown In Monocultures And Mixtures
- Grazing Resources Management
- Reliable And Cost Effective Legume Establishment In Black Speargrass Grazing Lands
- The Impact Of Soil Fertility And Legumes On The Yield And Persistence Of Buffel Grass
- Development Of Management Guidelines For Native Grass Pastures On The Central And Southern Tablelands Of NSW
- The Production And Management Of Annual Pasture Legumes In Ley Farming Systems Of South Australia
- The Role Of Lucerne In Developing The Agricultural Systems Of Eastern Gansu, China
- Establishment And Regeneration Of Annual Pasture Legumes In A Ley Farming System
- Managing Information
- The Impact Of Agricultural Information On Members Of Parliament.
- Strategies For Crop Information Management: Past And Future
- Issues Affecting Extension In The Dairy Industry
- Rapid Rural Appraisal Re Focuses Farming Systems Extension And Research In Southern Queensland
- An Analysis Of A Field Crop Agricultural Knowledge And Information System
- Farm Management Back To Basics
- A Framework For Comparing Crop And Pasture Rotations In North Western NSW
- Improving Wheat Profitability Through Farmer Initiated Groups
- The Role Of Extension Literature And Other Communication Media In Information Transfer
- Modelling And Decision Support
- An Economic Perspective On Pasture Research Priorities
- Decision Support For Dynamic Management On Extensive Beef Properties
- A Framework For Understanding The Long Term Value Of Pasture Leys In Cropping Systems
- Changing Grazing Systems With Experienced Local Graziers
- Competition Between A Crop And Undersown Pasture In A Water Limited Environment A Simulation Study
- Identifying Strategies To Minimise Climatic Risk For Grain Sorghum, Using The SORKAM Model
- A Framework For Research On Improved Management Of Agricultural Production Systems
- Seasonal Climate Forecasting In Crop Management
- Using A Model To Quantify Climatic Risk To Dryland Sunflower Production
- Pasturepak: An Electronic Pasture Adviser Built On Previous Results And Experience
- New Plants
- The Domestication Of Native Grasses For Pastoral Use
- Lotus In South Eastern Australia: Aspects Of Forage Quality And Persistence
- Domestication And Use Of Curly Mitchell Grass As A Sown Pasture
- Breeding Indian Mustard For Australian Conditions
- The Evaluation Of Linola As A New Oilseed Crop For Australia
- Development Of A New Mid Season Cultivar Of Subterranean Clover Conforming To A Predefined Ideotype
- New Technologies
- Mapping Soil Analysis Data In South East Australia
- Development And Commercial Use Of Landsat Derived Maps As An Aid To More Effective Use Of Fertilizer
- Airborne Video Imagery A Potential Tool For Monitoring Irrigated Cotton
- The Effect Of Landuse, Landform And Topographic Relief On Nocturnal Land Surface Temperature
- Plant Nutrition
- Response Of Pigeon Pea To Iron, Nitrogen And Two Rhizobium Strains
- An Alternative Hypothesis For The Nitrogen By Environment Interaction In Rice
- Potentially Mineralizable N A New Role In Predicting Soil N Supply
- Is Pasture Improvement Causing Soil Acidification In Northern NSW?
- Microarthropods And Nutrient Transfer From Pasture Litter
- Soil Stress
- Sustainability & Environment
- Productivity Of Oldman Saltbush In Relation To Rainfall And Economic Considerations
- Achieving The Potential Of The Pasture Resource In South West Victoria
- Traditional Management For Dryland Farming: Lessons For Modern Agronomic Systems Development
- Nitrogen Fixation In Mungbeans Expectation And Reality
- Incorporation Of Intensive Cropping Into Sustainable Systems In The Coal Valley, Tasmania
- Perceptions Of Sustainable Grazing Systems
- Practical And Economic Considerations Of Sustainable Agriculture In Summer Rainfall Areas Of Australia
- Applying Sustainability Through Farm Planning
- Sustainability Of Organic Broadacre Cropping Systems With Respect To Aspects Of Nutrient And Energy Flow
- Agriculture In The Wimmera; Looking Back And Planning Ahead
- Tillage And Soil Structure
- Direct Drilling With A Modified Combine Improves Root Growth, And Wheat Yields In Compaction Prone, Wind Erodible Soils
- Nitrogen Fertilisation Of Wheat Under Different Fallow Management Practices In Near South West Queensland
- Cereal Root Disease Constraints To Developing Conservation Tillage In Southern Australia
- Herbicide Carryover In Wheat Stubbles
- Spatial Variation In The Growth Of Lupin Roots In The Sandy Surface Of A Duplex Soil
- Effects Of Zero Tillage Practices On Soybean Yield And Soil Cover
- Alternative Practices And Plant Protection
- Conference Information
- Invited Addresses
- Specific Perspectives In Future Agriculture: Agricultural Sustainability
- The Past And Future Of Agriculture: Resolving Environmental Conflict
- Predicting And Shaping The Future: Managing Information
- Agriculture: Resources And Current Practices
- Managing Resources The Soil Resource: Erosion, Stubble Management And Catchment
- Cropping Systems Of The Temperate Summer Rainfall Region
- The Grazing Resource
- Socio Politics And Land Use Systems
- Initiatives In Land Use: Nutrients, Nutrient Cycling And Soil Acidity
- Initiatives In Land Use Soil Salinity
- Towards Sylvan Australian Farming Systems
- New Technologies
- Developments New Kinds Of Plants
- Protecting Yield – Chemically
- Protecting And Enhancing Yield, While Reducing Our Dependence On Synthetic Pesticides
- Agronomic Research Strategy And The Uncertain Challenge Of Sustainability
- Predicting And Shaping The Future: Models Looking Back And Planning Ahead
- The Future Is Not What It Was: Technological Perspectives On The Future Of Agriculture
- Concluding Review
- Crop Physiology: Some Recollections And Current Perceptions
- Poster Sessions
- Alternative Practices And Plant Protection
- The Impact Of Grass Control On Merino Lamb Production In The Cereal Belt Of South Australia
- Glyphosate Residues Persist In A Sandy Soil
- Integration Of Herbicide And Pasture Management For Long Term Control Of Vulpia
- Pasture Density Reduces The Establishment And Regeneration Of Vulpia
- Field Selection For Tolerance To Redlegged Earthmite And Lucerne Flea In Seedling Lucerne
- Integrated Screening For Resistance To Diseases And Pests In Lucerne Breeding
- Rampion Mignonette, A Mediterranean Weed New To Australia
- Breeding Phytophthora Resistant Chickpea
- Application Of Glyphosate To Conserve Annual Pasture For Grazing During Summer
- Grass Control In Pastures And Implications For Pasture Yields, Botanical Composition, Cereal Root Disease (take All) And Cereal Grain Yield
- Simulating Allelochemical Production From Plant Residues During Decomposition
- The Effect Of Broad Leaved Herbicides On The Pattern Of Water Use In Trifolium Subterraneum
- Cropping Systems
- Low Efficiency Of Wheat Production In The Victorian Wimmera In Above Average Seasons
- Improving Growth And Yield Of Wheat And Lupin Crops On Duplex Soils In Western Australia
- Late Planted Maize: A Source Of Low Productivity For Farmers In Subhumid Areas Of Zimbabwe
- Measuring The Economic Value Of Crop Research Proposals
- P 05
- P 06
- P 07
- P 08
- P 09
- P 10
- P 11
- P 12
- P 13
- Rotational Benefits Of Alternative Crops Can Be Affected By Their Sowing Time
- Grazing Resources Ecology
- Establishment Of Surface Sown Pastures In Wheat Stubble In North Western New South Wales
- Annual Medic Seed Reserves In Cereal Rotations
- Population Dynamics Of Volunteer Annual Grass Species As Affected By Spring Pasture Management
- Verano Stylo Seed Production In The Douglas Daly District, Northern Territory, Australia
- Aphids Can Alter The Pattern Of Seed Softening In Burr Medic
- Temperature Conditions Associated With The Autumn Softening Of Hard Seeds Of Burr Medic
- Subterranean Clover Seedling Survival In A Summer Rainfall Environment
- Effect Of Summer Rainfall On Seed Losses Of Two Forage Legumes, In The Wheatbelt Of Western Australia
- Effect Of Burial On Longevity Of Seeds Of Medicago Polymorpha Andtrifolium Subterraneum
- The Botanical Composition Of High Rainfall Pastures In South Western Victoria
- Annual Medics On Red Brown Earths
- Botanical Composition Of Pasture In South Western Victoria
- Using The Levy Point Quadrant To Assess Botanical Composition Of Dairy Pastures In The Adelaide Hills
- Time Of Planting Affects Seed Production Of Burr Medic
- Intake, In Vivo And In Vitro Digestibility’s, And Seed Survival Following Ingestion Of Annual Medic Pods By Sheep
- Effects Of Seed Size And Level Of Hard Seededness On Survival Of Medic Seeds In Whole Pods Fed To Sheep Or As Clean Seed Exposed In The Sheep Rumen
- The Significance Of Seed Size On Survival Of Some Annual Clover Pasture Species In South Australia
- Influence Of Defoliation Intensity And Time Of Final Defoliation On Seed Yield Of Paraggio Barrel Medic
- Predicting The Emergence Of Annual Pasture Legumes On Cereal Farms
- Effects Of Water Deficit During Vegetative Growth On Leaf Expansion Of Legumes
- Grazing Resources Management
- Perennial Ryegrass Improvement For Temperate Pastures In Australia
- Annual And Perennial Pastures React Differently To Autumn Winter Grazing Management
- National White Clover Improvement Programme The Plant Improvement Process
- The National White Clover Genetic Resource Centre
- Field Testing In The National White Clover Improvement Programme
- National White Clover Improvement Support Research
- Nodulation And Growth Of Serradella At Low Root Temperatures
- The Persistence Of Phalaris Under Grazing
- More Effective Rhizobial Inoculants Improve The Persistence Of Lucerne (medicago Sativa) In Western Australia
- The Effect Of Maturity On Proportion And Quality Of Morphological Components Of White Clover Grown Under Irrigation In Northern Victoria
- Effects Of Defoliation Management On The Productivity And Clover Content Of Irrigated Perennial Pastures
- Effect Of Time Of Removal Of Annual Grasses From Pastures On The Carryover Of Take All To Wheat
- Improved Production From Undersown Medics
- Pasture Consumption By Rabbits In The Central Highlands Of Victoria
- The Relationship Between Digestibility And Clover Content Of Dairy Pastures In The Adelaide Hills
- Managing Information
- A Message About Extension: Stimulate Not Saturate
- Haymaker
- Towards More Appropriate And Friendly Computerised Extension Tools Wiregrass Control Economics Advisor An Example
- Community Based Extension To Develop Productive Conservation
- A Land Management Manual For Waggamba Shire An Information Package Based On Land Resources
- Irrigation Scheduling And Crop Management System
- Modelling And Decision Support
- New Plants
- An Evaluation Of Lines Of Trifolium Balansae For Late Maturity
- Development And Potential Of Holdfast Phalaris
- Breeding Narbon Beans To Establish A New Crop For South Eastern Australia
- Medicago Orbicularis (l.) Bart. A Potential New Medic For The Cereal Belt Of Southern Australia
- Breeding And Selection Of Improved Cultivars Of Trifolium Subterraneum Ssp. Brachycalycinum
- Ambiphotoperiodic Flowering Response Of Kenaf (cv. Guatemala 4)
- Fibre Flax, An Old And A New Crop In North Western Germany
- New Technologies
- Improving Fertiliser Use Efficiency By Deep Banding
- Effect Of Water Injection And Seed Soaking On Establishment Of Sunflower In A Vertisol
- Soybean Desiccation Diquat, Ethephon And Windrowing
- Effect Of Size Grading On Lupin Productivity
- Operation Undersow
- The Importance Of Low Velocity Wind Turbulence In Reducing The Rate Of Frost Development
- Pasture
- The Wool Production Zones Of Australia Ii. Perennial Pasture Zone Productivity & Pasture Problems
- The Wool Production Zones Of Australia Iii. Productivity & Management Of The Rangelands
- The Pasture Resource Base
- Management For Pasture Establishment
- The Management Of Nutrients For Pastures
- Grazing For Pasture Management In New Zealand
- Grazing For Pasture Management In The High Rainfall, Perennial Pasture Zone Of Australia
- Grazing For Pasture Management Annual Pasture Zone
- Grazing For Pasture Management In The Rangelands
- Animal And Pasture Maintenance, A Partnership
- Pasture Management At The Farm Level
- The Wool Production Zones Of Australia In. The Annual Pasture Zone Productivity & Pasture Problems
- Plant Nutrition
- Effects Of Nitrogen Application On The Light Interception And Radiation Use Efficiency Of Sunflower, Helianthus Annuus L.
- Mineralisation Of Soil Organic Nitrogen Under Long Term Rotations
- Cereal Grain Yield Response To Pasture Legume Nitrogen
- High Concentrations Of Carbon Dioxide Decrease Critical Nitrate And Total Nitrogen Concentrations In Wheat
- Relationships Of Petiolar Nitrate Concentrations With Growth And Yield Of Sunflower
- Use Of Plant Analysis To Assess The Sulfur Status Of Field Grown Subterranean Clover
- Penalties Of Ceasing Phosphorus And Sulphur Applications, To Pastures On The North West Slopes Of NSW
- The Relative Tolerance Of Balansa And Sub Clovers, Murex Medic And Lucerne To Aluminium And Manganese
- The Use Of S" As An Index Of The Sulfur Status Of Plants
- Response Of Wheat To Four Nitrogen Management Systems
- Effect Of Nitrogen Fertilizer On The Yield And Grain Nitrogenconcentration Of Malting Barley
- Lime Response Of Various Native Grasses
- Effect Of Nitrogen On Yield And Protein Content Of Oats
- Fertiliser Key To Profitable And Sustainable Pasture Lands, Northern Slopes And Plains, Central Slopes, Central West Plains And Upper Hunter
- Soil Potassium Decline In Western Australian Agricultural Soils
- Pasture And Cereal Responses To Applied Lime In Central NSW
- Effects Of Aeration And Nitrogen Supply On Cotton Growth In Solution Culture
- Soil Stress
- Waterlogging In A 450 Mm Rainfall Zone In South Australia?
- Drought Tolerance Of Wheat Seedlings
- Re Use Of Saline Water On Tomatoes: An Approach To Managing Salinity And Increasing Profit
- Effect Of Timing Of Waterlogging On Survival Of Chickpeas (cicer Arietinum L.)
- Water Management For Irrigated Pasture And Salinity Control In South West Western Australia
- Productivity And Properties Of Red And Grey Gradational Soil Types In The Victorian Wimmera
- Sustainability & Environment
- VAMS Of Temperate Pastures In The High Rainfall Zone
- Conservation Cropping On The Atherton Tableland Krasnozems
- Reduced Tillage In Tobacco
- Halophyte Agronomy On Dryland Saline Areas In South West Victorian High Rainfall Districts
- Comparison Of The Effects Of Synthetic, Mineral And Organic Fertilisers On Barley Production
- A Comparison Of Farming Systems
- Is Continuous Cropping Sustainable On Red Duplex Soils?
- Crop Rotation For Broadacre Organic Cropping Systems In South East Australia
- Extending Principles Of Sustainable Agriculture
- Alternative Practices And Plant Protection
- Concurrent Sessions
- 1993 7th AAC
- Concurrent Sessions
- Crop Nutrition
- Potential Residual Value Of Nitrogen Fertilisers In Central Queensland 2
- Wheat Yield Targets, And Water And Nitrogen Use Efficiency In Northern New South Wales 3
- Wheat Responses To Nitrogen And Phosphorus Application In The Semi Arid Environment Of Central Queensland 4
- Critical Level For Zinc Deficiency Of Field Grown Wheat 5
- Management Factors That Affect Cadmium Uptake By Wheat Grain 6
- The Effect Of Nitrogen On The Yield And Selected Aspects Of Quality Of Two Varieties Of Malting Barley Grown Under Contrasting Conditions Of Water Availability 1
- Extension, Surveys And Marketing
- Farmer Attitudes Towards Pastures In The Cereal Livestock Zone Of South Australia 2
- Participatory Design Of An On Farm Agronomic Research Program In Central Queensland 3
- Strategies Used For Marketing And Promoting German Agricultural Products 4
- Market Research For Decision Support For Dryland Crop Production In Central Queensland: A Preliminary Report 5
- Integrated Marketing Of Extension Messages Lifts Adoption 6
- Digging Up The Past 1
- Grain Legumes And Oilseeds
- Modelling
- Assessment Of Risk Associated With Climate Prediction In Management Of Wheat In North Eastern Australia 2
- Climatic Risk For Sweet Corn Production In A Cool, Variable Climate 3
- Potential Yield Of Peanuts In North Eastern Australia 4
- Modelling Rice Growth Under Water Limiting Conditions 5
- Scenarios Of Climate Change In The Australian Region Due To The Enhanced Greenhouse Effect 1
- Can Profitable Cropping Systems Be Developed In The Semi Arid Tropics Of Northern Australia? 2
- Analysis For Optimal Water Use In Grain Cropping Systems Of North Eastern Australia 3
- Agricultural Land Capability Assessment Using An Expert System 4
- APSIM: The Agricultural Production System Simulator Its Role And Structure 1
- Pastures : Composition
- Seasonal Pattern Of Growth Of Subterranean Clover Grass Pasture 2
- Grass Control In Medic Pastures In Semi Arid Ley Farming Systems 3
- The Impact Of Herbicides On The Production, Composition And Quality Of Annual Pasture At Kapunda, South Australia 4
- Grass Control In Pastures: Implications For Livestock 5
- Control Of Onion Grass (romulea Spp.) Prior To Resowing Permanent Pastures 6
- Manipulating Pasture Composition A Matter Of Time, Dollars And Sense 1
- Pasture Management : Establishment And Ecology
- Current Management Of Pastures On Properties On The Tablelands Of NSW 2
- The Significance Of Cluster Clover (trifolium Glomeratum L.) In A Subterranean Clover Perennial Ryegrass Pasture Under Two Constrasting Management Systems. 3
- Regeneration And Recruitment Of Curly Mitchell Grass (astrebla Lappacea) In Degraded Pastures In North Western New South Wales 4
- The Use Of Open Rooted Seedlings In Farm Tree Plantations 5
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic 1
- Pastures Breeding And Selection
- Developing Well Adapted Trifolium Subterraneum Ssp. Brachycalycinum Cultivars 2
- Balansa And Persian Clover Lines Outproduce Registered Cultivars, And Subterranean Clover And Medics, In A 400 Mm Annual Rainfall Zone In Western Australia. 3
- Evaluation On Growth And Production Of 13 Provenances Of Gliricidia Septum 4
- Maximising Forage Production From Saltbushes Grown On Salt Affected Agricultural Land 5
- Selection Of Localised Strains Of Trifolium Subterraneum Var. Brachycalycinum Adapted To A Summer Rainfall Environment 1
- Pastures And Fodder Crops : Utilisation
- The Nutritive Value Of Medic Pods Following Treatment With Sodium Hydroxide Or Supplementation With Molasses 2
- Turnips, The Superior Fodder Crop In South Western Victoria 3
- Biodynamic Milk Producing Systems 4
- New Market Alternatives For North Australian Beef 5
- Spring Grazing Management Of Medic Pastures To Optimise Productivity Of Herbage And Seed 1
- Plant Physiology
- Responses Of Barley, Wheat And Maize To Drought In A Cool Temperate Climate 2
- Influence Of Water Stress On Leaf Death Among Rice Lines: Comparison Between Glasshouse And Field 3
- Tolerence Of Temperature Stress In Phaseolus Vulgaris. 4
- Leaf Nitrogen Profiles In Sunflower Canopies During Grain Filling 5
- Apsim: The Agricultural Production System Simulator Its Role And Structure 6
- Can Profitable Cropping Systems Be Developed In The Semi Arid Tropics Of Northern Australia? 7
- Analysis For Optimal Water Use In Grain Cropping Systems Of North Eastern Australia 8
- Agricultural Land Capability Assessment Using An Expert System 9
- The Nature And Agronomic Implications Of A Juvenile Stage In Barley 1
- Managing The Soil Resource
- No Tillage Wheat Production In Northern NSW 2
- Effects Of Surface Soil Mixing After Long Term Disc, Blade And Zero Tillage On Sorghum Production In Central Queensland 3
- Soil Conservation In The Paddock Keeping A Full Plate Into The Future Effects Of Farming Practices On Red Soils In Central West NSW. 4
- Effect Of Land Management Systems On Rainfall Infiltration Capacity And The Stability Of Red Brown Earths 5
- The Influence Of Fertiliser On The Abundance And Diversity Of Earthworms In Pastures In Western Victoria 6
- Sustainability Healthy Paddocks Lead To Full Plates 1
- Summer Crops
- Use Of A Simple Physiological Model To Analyse Yield Variation In Phaseolus Vulgaris Genotypes 2
- Variation In Root Characteristics, And Their Association With Water Uptake And Drought Tolerance In Four Peanut Cultivars 3
- Knowledge Of Stand Variability Aids Interpretation Of Hand Thinned Experiments 4
- Saturated Soil Culture: A New Concept For Irrigated Rice Production In Tropical Australia 5
- Mungbean Development In The Northern Territory 1
- Weeds And Weed Control
- Spread Of Herbicide Resistance Through Seed Retail Outlets 2
- Biological Activity Of Barley Secondary Metabolites 4
- The Effect Of Soil On The Phytotoxicity Of Residues Of Vulpia Myuros 5
- Herbicides And Equipment To Improve Pastures Infested With Cape Tulip (homeria Spp.) 6
- On The Merits Of Setting Targets For Pesticide Use 1
- Winter Cereals And Rotations
- Crop Nutrition
- Conference Information
- Invited Addresses
- Grain Legumes: Farming From Paddock To Plate
- Production Of Quality Wool A Case Study
- Future Markets For Australian Agricultural Produce
- Utilisation Of Joint Action Groups To Achieve Export Impact
- Research And The Restructuring Of Australian Agriculture: Transforming Comparative Advantage Into A Competitive Advantage
- Global Perspective On Population, Resources And Agricultural Production
- Agricultural Science And Technology Meeting The Challenge
- Durum Wheat A Specific Market
- Elite Lambs For Meat Production
- Poster Sessions
- Crop Nutrition
- Response Of Barley Varieties To Nitrogen Fertiliser Under Contrasting Water Availability 2
- Cultivar Differences In Responses To N Fertilizer In Malting Barley 3
- Effect Of Time Of Application Of Nitrogen Fertilizer On Dry Matter And Nitrogen Partitioning In Wheat 4
- Response Of Wheat To Time Of Nitrogen Fertilizer Application In A Cool Temperate Climate 5
- Comparison Of The Effects Of Synthetic, Mineral And Organic Fertilisers On Medic Production 6
- Efficient Fertiliser Management Of Wheat Crops In The High Rainfall Western Districts Of Victoria. 8
- Grain Yield, Nitrogen Uptake And Sustainable Wheat Production: Are Varieties Equal? 9
- Some Zinc Enriched Fertilizers Are Not Effective At Correcting Zinc Deficiency 10
- The Effect Of Nitrogen On The Yield And Protein Concentration Of Soft Wheat 1
- Farm Surveys And Demonstrations
- Crop Rotations Used In The Cotton Industry 2
- How Much Feed Is In The Paddock? 3
- What Layout And Lettering Will Lift Readership ? 4
- Improving Annual Pastures In The South Australian Cereal Belt 5
- The Pasture Legume Status Of Cereal Farms In The Central Region Of South Australia 6
- The Adoption Of Pasture Management Practices In The Murray Mallee Of South Australia 7
- Farmer Managed, On Farm Demonstration Of The Impact Of Pasture Technology On Crop And Livestock Production 1
- Germination And Establishment
- Effect Of Nutrient And Fungicide Seed Coating On The Emergence And Dry Matter Production Of Tall Fescue And Red Clover 2
- Effects Of Defoliation On Pod Set And Seed Yield In A Range Of Medic Cultivars 3
- Comparison Of Resowing Techniques For Perennial Grass/subclover Pastures In The Mt. Lofty Ranges 4
- Predicting Annual Medic Emergence In The Murray Mallee Of South Australia 6
- Pasture Manipulation To Improve Crop Pasture Rotations In Southern Australia 7
- Invertebrate Pests Associated With Annual Medic Pastures In South Australia 8
- Herbicide Effectiveness And Subsequent Pasture Establishment In Direct Drilled White Clover Pastures 9
- The Effect Of Various P And N Fertilisers And Lime On The Establishment Of Direct Drilled Pasture 10
- Grain Legumes And Oilseeds
- Nitrogen And Sulfur Affect Canola Seed Yield And Oil Percentage 2
- The Effect Of Seed Composition On Seedling Vigour In Some Pea Cultivars 3
- The Effect Of Seed Size, Seed Protein And Genotype On Seedling Vigour In Some Grain Legumes 4
- Improved X Ray Spectrometric Method For Determining Glucosinolates 5
- Factors Influencing Yield Of Gungurru Narrow Leafed Lupins From Different Sources In Western Australia 6
- The Effect Of Sowing Time And Plant Density On The Grain Yield And Dry Matter Of Fara Bean (Vicia Faba L.) 1
- Modelling
- Long Term Agronomic Trials In Australia 2
- An Integrated Decision Support System For Pasture Improvement 3
- A Framework For Developing Agricultural Production System Models 4
- APSIM + GrazPlan: Versatile Software For Simulating Grain Grazing Systems 5
- Assessing The Performance Of Maize (zea Ma Ys) Cowpea (vigna Unguiculata) Intercrop Under Variable Soil And Climate Conditions In The Tropics 1
- Pastures : Cultivars And Selection
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic
- Effects Of Sowing Date And Sowing Rate On Seed Yield Of Barrel Medic
- Plant Selection Of Perennial Pasture Legumes And Cocksfoot For A Niche Environment 2
- Selection Of A Triplex Species For Salt Pan Rehabilitation And Fodder Production In The Lockyer Valley 3
- A Collection Of Naturalized Annual Pasture Legumes In The Semi Arid Cereal Livestock Zone Of South Australia 4
- The Growth And Development Of Exotic Medicago Spy. At Roseworthy, South Australia 5
- Incidence And Agronomic Significance Of Endophyte In Perennial Ryegrass Seed Lines 6
- Caliph A New Barrel Medic Cultivar For Southern Australia 1
- Pastures : Grazing
- Height Dry Matter Relationships In Subterranean Clover Grass Pasture 2
- Grazing Behaviour Of Race Horses On Perennial Ryegrass 3
- Comparision Of Crop Residues For Ruminant Feed 4
- Variation In Feeding Value Of Senescent Annual Ryegrass 5
- The Comparative Effect On Medicago Orbicularis Seed Reserves Of Grazing Dry Residues 6
- Influence Of Maturity On The Nutritive Value Of Red Clover Grown Under Irrigation In Northern Victoria 7
- Nutritive Value Of Cereal Straws And Grain Legume Residues 8
- The Potential For Lot Feeding Of Sheep On Cereal Farms To Protect Medic Seed Reserves And Surface Soil. 1
- Pastures : Rhizobia
- Effects Of 2,4 Dr (amine Salt) On The Growth, Nodulation And Nitrogen Fixation Of Annual Medics 2
- Lucerne Decline The Role Of Soil Acidification And Herbicides 3
- The Effect Of Glean Residues On Growth And Nodulation Of Medic In An Alkaline Dark Brown Sandy Loam 4
- Survival Of Rhizobium Trifolii With Seed Dressing Fungicides 1
- Plant Physiology
- Rangeland Grass Growth With Nitrogen Limitation Is Increased By High Levels Of Atmospheric Carbon Dioxide 2
- Dynamics Of Rooting Patterns For Wheat Cultivars Of Different Cycle Lengths. 3
- Number Of Grains In Modern And Old Cultivars Of Wheat, Triticum Aestivum, Grown Under Preanthesis Shading. 4
- The Use Of Wild Wheats To Increase Potential Grain Number Per Head In Hexaploid Wheat 1
- Soil Constraints
- Water Use
- Potential Water Use Of Phalaris, Cocksfoot, Lucerne And Birdsfoot Trefoil Cultivars With Varying Seasonal Growth Patterns 2
- Comparative Water Use Of Yellow Serradella, Volunteer Pasture And Wheat On A Sandplain Soil In Western Australia 3
- Effects Of Soil Management And Irrigation Method On The Yield And Water Use Efficiency Of Soybean 4
- Irrigation Frequency For Perennial Pastures In The Murrumbidgee Valley 1
- Weeds And Weed Control
- Should Farmers Be Concerned About The Spread Of The Pasture Weed Water Dropwort, Oen Anthe Pimpinelloides? 2
- Broad Leaved Weed Control In Annual Medic Pastures 3
- Sulfonylureas And Annual Medic Regeneration On High PH Soils 4
- Preliminary Observations Of Pasture Legume Herbicide Tolerance 5
- Integrated Weed Management The Role Of Competitive Cultivars 6
- Rampion Mignonette Current Research Into A New Weed 7
- Competitive Effects Of Crop Species On Skeleton Weed Populations 8
- Calepina, Calepina Irregularis And Turkish Mustard, Malcolmia Africana Two Mediterranean Weeds New To Australia. 1
- Winter Cereals
- Crop Pasture Rotations: Are They The Key To Better Crops In North Western NSW? 2
- Crop Establishment Improvement From Narrow Spear Points In South Australia 3
- Plant Sampling Methods For Seed Drill Research 4
- The Effect Of Three Medic Based Pastures On Take All And Root Lesion Nematode Levels 5
- The Effect Of MCPA And Dicamba On Take All Disease And Growth Of Wheat Seedlings 6
- The Potential Of Australian Wheat For German Egg Noodles (spatzle) 7
- Timely Seeding Is A Re Think Necessary In South Australia? 8
- Triticale: Its Place In Agriculture, And Marketing 1
- Crop Nutrition
- Concurrent Sessions
- 1996 8th AAC
- Contributed Papers
- Farmers’ Risk Perspective On Adoption Of Legume Crops
- VARIATION OF WHEAT AND SUBTERRANEAN CLOVER CULTIVARS IN SUSCEPTIBILITY TO VULPIA RESIDUE PHYTOTOXICITY
- TOPCROP Integrated Marketing Meets Extension
- IMPROVED PASTURES AND THEIR EFFECTS ON SUBSEQUENT CROPS
- MaNage Rice: DECISION SUPPORT FOR TACTICAL CROP MANAGEMENT
- Sustainable Resource Use And The Evaluation Of Agronomy R&D
- THE EFFECT OF NON LETHAL WATER DEFICITS DURING ESTABLISHMENT ON THE GROWTH OF WHEAT CROPS
- UTILISATION OF NITROGEN FROM PLANT RESIDUES AND FERTILISERS BY WHEAT IN CENTRAL QUEENSLAND FARMING SYSTEMS
- EFFECTS OF SURFACE SOIL MIXING AFTER LONG TERM ZERO TILLAGE ON SOIL NUTRIENT DISTRIBUTION AND WHEAT PRODUCTION
- EFFECTS OF TILLAGE PRACTICE AND LEY PASTURE ON SOYBEAN PRODUCTIVITY ON A DEGRADED FERROSOL
- Quantifying The Yield Density Relationship For Narrow Leafed Lupin (lupinus Angustifolius) In Tasmania
- When Adoption Theory Fails
- ARE DISTRICT NITROGEN RECIPES OBSOLETE?
- An Integrated Program To Improve Anthracnose Resistance In Stylosanthes A Review
- ESTABLISHMENT OF SURFACE SOWN PASTURES ON UNPLOUGHED INFERTILE SOIL
- FARMER DISCUSSION GROUPS PAST, PRESENT AND FUTURE
- Differences In Radiation Use Efficiency Among Lines In A Tropical Maize Population
- SOIL MANAGEMENT STRATEGIES FOR INCREASED WINTER PASTURE PRODUCTION ON DRYLAND DAIRY FARMS
- AGRONOMY AND FARM VIABILITY
- THE POTENTIAL OF UPGRADED PERENNIAL PASTURES TO REDUCE GROUNDWATER RECHARGE IN SOUTHERN VICTORIA
- Measuring The Effects Of Landuse And Land Management On River Water Quality
- INCREASING COMMITMENT TO PASTURES IS THE KEY TO IMPROVING FARM PRODUCTIVITY IN THE EAST GIPPSLAND WOOL INDUSTRY
- SURFACE SOIL NUTRIENT DISTRIBUTION FOLLOWING ZERO TILLAGE AND TRADITIONAL TILLAGE MANAGEMENT
- CAUGHT IN THE WEB: USE OF THE INTERNET IN AGRICULTURAL R,D&E
- RE ENGINEERING AGRICULTURAL R,D&E TO SUPPORT MANAGEMENT DECISION MAKING: PROBLEMS AND PROSPECTS
- BELOW GROUND INPUTS OF CARBON BY CROPS AND PASTURES
- EFFECTS OF FALLOW MANAGEMENT AND CROPPING SEQUENCES ON PRODUCTION AND RESIDUAL N UPTAKE IN WHEAT
- SUSTAINABILITY AND PROFITABILITY: A CASE STUDY FROM THE WARRA EXPERIMENT
- DEVELOPING WELL ADAPTED EARLY TO MIDSEASON CULTIVARS OF TRIFOLIUM SUBTERRANEUM SSP. BRACHYCALYCINUM AN UPDATE
- PRELIMINARY INVESTIGATION OF LOCALLY ADAPTED ECOTYPES OF TRIFOLIUM SUBTERRANEUM SSP. BRACHYCALYCINUM CV. CLARE IN SOUTH AUSTRALIA
- FACTORS INVOLVED IN ANNUAL MEDIC DECLINE SYNDROME IN THE MURRAY MALLEE, SOUTH AUSTRALIA
- EFFECT OF PHOSPHORUS RATE AND PLACEMENT ON EARLY GROWTH AND NODULATION OF FIELD PEAS (PISUM SATIVUM L. Cv ALMA)
- WHEAT PRODUCTION AND SOIL CHEMICAL PROPERTIES OF ORGANIC AND CONVENTIONAL PAIRED SITES IN WESTERN AUSTRALIA
- RARE EARTH ELEMENTS AND PLANT GROWTH
- HOWWET? A TOOL FOR PREDICTING FALLOW WATER STORAGE.
- PERSISTENCE OF TEMPERATE PASTURES IN A SUBTROPICAL ENVIRONMENT
- MANAGING NITROGEN IN CROPPING SYSTEMS FOLLOWING LEGUME LEYS
- THE EFFECT OF SOIL AMMONIUM CONCENTRATION AND OSMOTIC PRESSURE ON SEEDLING EMERGENCE
- THE SOCIAL IMPACT OF COMPUTER NETWORKING ON THE RESTRUCTURING OF AN EXTENSION SERVICE
- ALTERNATIVES TO THE SOFTWARE PRODUCT MENTALITY IN EXTENSION
- INTERCROPPING WHEAT, OATS AND BARLEY INTO LUCERNE IN VICTORIA
- SEWAGE SLUDGE: RESOURCE OR POLLUTANT
- THE EFFECT OF NITRATE CONCENTRATION ON THE GROWTH AND NODULATION OF SEVERAL ANNUAL SPECIES OF MEDICAGO
- THE EFFECT OF ROW SPACING AND SEEDING RATE ON CHICKPEA YIELD IN NORTHERN NEW SOUTH WALES
- PRIME HARD WHEAT AND MALTING BARLEY HOW DOES PROFILE NITROGEN INFLUENCE THE FARMER’S CHOICE?
- FARMERS, ADVISERS AND RESEARCHERS LEARNING TOGETHER BETTER MANAGEMENT OF CROPS AND CROPLANDS
- A SUCCESSFUL PASTURE SOIL NUTRITION PROGRAM
- Nutrient Threat To Sustainable Agriculture
- MAINTAINING WHITE CLOVER (TRIFOLIUM REPENS) IN A KIKUYU (PENNISETUM CLANDESTINUM) PASTURE IN THE SUBTROPICS
- HIGH CONCENTRATIONS OF SOIL SOLUTION PHOSPHORUS IN WIMMERA SOILS
- SYSTEMS MODELLING FRAMEWORK FOR CATCHMENT SCALE SALINITY MANAGEMENT
- ROTATION WITH CANOLA AND OPTIMISING WHEAT YIELD IN NORTH EAST VICTORIA
- SOIL TESTING IN THE PADDOCK: AN EVALUATION OF A FIELD TEST KIT
- A CASE STUDY IN DECISION SUPPORT INFORMATION: A MANUAL TITLED MAKING BETTER DECISIONS FOR YOUR PROPERTY
- OPPORTUNITY CROPPING ON THE LIVERPOOL PLAINS: A COMPARISON OF RISK ASSESSMENT BY FARMERS AND SIMULATION MODELS
- ADAPTING A BARLEY GROWTH MODEL TO PREDICT GRAIN PROTEIN CONCENTRATION FOR DIFFERENT WATER AND NITROGEN AVAILABILITIES
- Wheat Breeding In Germany, Assisted By Modern Equipment And Aspects Of Tomorrow
- HALOSULFURON METHYL AND IMAZETHAPYR For The SELECTIVE CONTROL OF KYLLINGA (CYPERUS BREVIFOLIUS ROTTB.) IN IRRIGATED PASTURE
- MANAGING THE PASTORAL LANDSCAPE: ADDING A SPATIAL DIMENSION?
- NITROGEN FIXATION IN CHICKPEA AS AFFECTED BY PLANTING TIME AND TILLAGE PRACTICE
- The Use Of Crop Monitoring For Problem Analysis And Decision Making In Wheat Crops
- DO YOUR OWN RESEARCH IN THE NURSERY INDUSTRY
- ENVIRONMENTAL VANDALISM? TOWARDS AN OBJECTIVE ASSESSMENT OF WEED STATUS
- RAISING SOYBEAN YIELD THROUGH APPLICATION OF CROP PHYSIOLOGY TO AGRONOMY AND BREEDING
- A NEW MODEL OF SPRING WHEAT PHENOLOGICAL RESPONSE TO TEMPERATURE AND DAYLENGTH
- MONITORING IMPROVEMENTS IN SEED PRODUCTION AND SEEDLING VIGOUR OF GREATER LOTUS (LOTUS PEDUNCULATUS Cav.)
- GRAIN OATS A POOR BREAK CROP FOR WHEAT
- Suppression Of Soil Borne Cereal Pathogens And Inhibition Of Wheat Germination By Mustard Seed Meal
- STRATEGIC Initiatives To Identify And Meet Market Opportunities In The AUSTRALIAN Grains Industry
- ANNUAL MEDIC CULTIVAR MIXTURES IN SEMI ARID FARMING SYSTEMS
- SOIL NITROGEN DECISION AID PUTS FARMERS IN CONTROL
- Improved Variety Testing And RELEASE Procedures In The AUSTRALIAN Grains Industry
- THE INFLUENCE OF ALLEY FARMING AND INTERCROPPING ON THE YIELD OF LUPINS
- DEVELOPMENT AND APPLICATION OF RFLP AND STS IN STYLOSANTHES
- Adaptation Of Faba Bean To Mediterranean EnvironmentS Of Western AUstralia
- PROMISING NEW GRAIN LEGUMES FOR WESTERN AUSTRALIA
- EVOLUTION AND ACCOUNTABILITY IN THE GRDC
- METHODS TO ASSESS SOIL CHEMICAL PROPERTIES AND THEIR RELATIONSHIP WITH FIELD MEASUREMENTS OF INFILTRATION AND EROSION.
- MANAGING THE NITROGEN NUTRITION OF WHEAT IN NORTHERN NEW SOUTH WALES USING NITROGEN BUDGETS
- CARRYOVER OF SOIL NITRATE N FOLLOWING FERTILISER APPLICATION TO WHEAT IN NORTHERN NEW SOUTH WALES
- DEVELOPING QUALITY WHEAT VARIETIES FOR THE MARKET
- AN INTEGRATED APPROACH TO LINK FARMING SYSTEMS RESEARCH, EXTENSION CO LEARNING AND SIMULATION MODELLING
- Integrated Weed Management For The Control Of Herbicide Resistant Annual Ryegrass
- AN INTEGRATED APPROACH TO EVALUATION OF RESEARCH OPPORTUNITIES IN THE QUEENSLAND SUGAR
- 425meyer
- EFFECTS OF SEWAGE SLUDGE ON PASTURE PRODUCTION AND SHEEP PERFORMANCE
- IMPROVEMENTS IN R, D & E ARE ON GOING
- PHENOLOGY AND GRAIN YIELD OF FIELD PEA IN THE NORTHERN CROPPING BELT
- THE POTENTIAL FOR GROWING SUGARBEET IN THE MACKAY AND BURDEKIN SUGARCANE AREAS
- THE FITZROY RIVER CATCHMENT: AN ASSESSMENT OF THE CONDITION OF THE RIVERINE SYSTEM
- BREEDING NON TOXIC PHALARIS (PHALARIS AQUATICA L.)
- APSIM SoilWat And SoilN: Validation Against Observed Datafor A Cracking Clay Soil
- Modelling Changes In Soil Microbial Biomass In Response To Added Crop Residues
- ESTABLISHMENT OF AN OPTIMAL GRAZING TIME OF KIKUYU PASTURES FOR DAIRY COWS
- MODELS AND DECISION SUPPORT: BRIDGING THE MODEL GAP
- INTERPRETATION AND VALUE OF SOIL NITRATE NITROGEN AT DEPTH
- APPLICATIONS FOR A NEW TECHNOLOGY: COMPACT WEATHER RECORDING SYSTEM
- DO MELILOTUS SPECIES HAVE A ROLE FOR SALINE AREAS IN AUSTRALIA?
- CHEMICAL FALLOWING AND MANIPULATION OF PASTURE ALONG WITH EARLY SEEDING GIVE GREATER YIELDS AND WATER USE OF WHEAT ON A SANDY CLAY LOAM SOIL
- The Effect Of Non Selective Herbicides On Pasture And Animal Production
- TRENDS IN CROP PRODUCTION AND SOIL PROPERTIES IN A LONG TERM TRIAL AT TARLEE, SOUTH AUSTRALIA
- IMPROVING THE IRRIGATION EFFICIENCY OF BURDEKIN CANEGROWERS
- THE AUSTRALIAN GERMPLASM OF PINK FLOWERING SERRADELLA SPECIES
- Comparisons Of New Medicago Spp. In South Australia
- INFORMATION NEEDS IN AGRONOMY ON THE DARLING DOWNS, QUEENSLAND
- LUCERNE/VEGETABLE ALLEY CROPPING TO CONTROL LEACHING
- PROFITABLE USE OF NITROGEN FERTILISER TO SUSTAIN PRODUCTION WHERE RAINFALL IS UNRELIABLE
- OPERATION UNDERSOW: IMPROVING PERENNIAL PASTURE ESTABLISHMENT UNDER A CEREAL CROP
- FACTORS LIMITING WHEAT YIELDS UNDER ZERO TILLAGEIN SOUTHERN QUEENSLAND
- Responding To Increased Demands For Accountability The Role Of Benefit Cost Analysis In Research Investment Decisions
- A MULTI MEDIA EDUCATIONAL PROGRAM IN PASTURE ASSESSMENT AND MANAGEMENT
- VEGETATIVE BARRIER WITH VETIVER GRASS: AN ALTERNATIVE TO CONVENTIONAL SOIL AND WATER CONSERVATION SYSTEM
- ADAPTATION OF COOL SEASON PULSES TOWATER LIMITED ENVIRONMENTS
- Computer Simulation Of The Effects Of Cropping Rotations And Fallow Management On Solute Movement
- Simulation Of Cereal Legume Rotations Using APSIM
- HAYING OFF IN WHEAT: ENDURING MYTH OR CURRENT PROBLEM?
- PHYSIOLOGICAL RESPONSES OF SIX SPRING WHEAT VARIETIES TO NITROGEN FERTILISER
- ASSESSING THE ENVIRONMENTAL IMPACTS OF THE GRAINS INDUSTRY: AN OVERVIEW
- DEPLETION AND RECHARGE OF AVAILABLE SOIL WATER UNDER THREE PASTURE LEYS AND CONTINUOUS WHEAT AT WARRA IN SOUTHERN QUEENSLAND
- CHANGES IN SOIL PHYSICAL PROPERTIES AND SOIL ORGANIC CARBON FRACTIONS WITH CROPPING ON A RED BROWN EARTH SOIL
- Identifying Persistent Lucernes For Dryland Pastures
- PEA PHENOLOGY RESPONSES TO TEMPERATURE AND PHOTOPERIOD
- ECONOMICS OF PASTURE ROTATIONS IN QUEENSLAND WHEAT AREAS
- FARMING SYSTEMS THE NEED FOR ECONOMICS AND INTEGRATION
- RUMEN FUNGAL Β GLUCANASE AND XYLANASE GENES: POTENTIAL FORGENETICALLY ENGINEERED CEREAL CROPS
- TRITIUM CONTENT OF SHALLOW GROUNDWATERS IN THELIVERPOOL PLAINS CATCHMENT
- Conference Information
- Invited Papers
- Poster Papers
- Field Application Of Vulpia Phytotoxicity Management:A Case Study
- ANHYDROUS AMMONIA APPLIED AS COLDFLO™ TO WHEAT
- EFFECTS OF CHILLING TEMPERATURES ON PHOTOSYNTHETIC RATE IN AUSTRALIAN PEANUT VARIETIES
- AN EXAMINATION OF THE REASONS FOR VARIATION AMONG CROP SPECIES IN THE UTILITY OF SOIL P TESTS YIELD RESPONSES ON A FERROSOL
- STRATEGIES TO REDUCE CADMIUM ACCUMULATION IN PEANUTS
- PREDICTING GROWTH OF WHEAT FOLLOWING PASTURE
- GRAZING MANAGEMENT OF LOTUS: A PARTICIPATORY APPROACH
- IMPROVEMENT OF KANGAROO VALLEY PERENNIAL RYEGRASS
- EFFECT OF DIFFERENT INTENSITIES OF DROUGHT AND DEFOLIATION UPON THE MORTALITY OF PERENNIAL GRASSES
- COMPENSATION IN SOYBEANS DUE TO ARTIFICIAL POD REMOVAL, A PREDICTOR OF COMPENSATION DUE TO INSECT DAMAGE
- A Chickpea Cultivar X Population X Row Space StudyIn Southern QUEENSLAND
- PADFERT 2
- THE EFFECT OF CROP SEQUENCE AND TILLAGE PRACTICE ON SOIL LOSS AND RUNOFF IN A SEMI ARID ENVIRONMENT
- FABA BEANS NEED HIGHER SEEDING RATES IN HIGH RAINFALL AREAS
- RISKS TO STYLOSANTHES CULTIVARS FROM ANTHRACNOSE DISEASE
- WATER QUALITY IN RUNOFF FROM NATURAL RAINFALL FROM DIFFERENT SURFACE MANAGEMENT TREATMENTS ON A MIAMIAN SOILIN CENTRAL OHIO, USA
- CONSEQUENCES OF SOIL STRUCTURE DECLINE ON CROPPING IN SOUTH EAST QUEENSLAND
- COMPARING ROTATION CROPS FOR SUSTAINABLE COTTON PRODUCTIONIN THE MACQUARIE VALLEY OF NEW SOUTH WALES
- GLYPHOSATE: A RE APPRAISAL OF THE THREAT TO CROP PLANTS
- Improving Nitrogen Application Decisions For Barley
- MEASURING PLANT AVAILABLE SOIL WATER STORAGE CAPACITY IN NORTHERN AUSTRALIA
- SOIL SAMPLING TECHNIQUES FOR MONITORING WATER AND NITROGEN
- HOST SPECIFICITY OF THE ANNUAL MEDIC RHIZOBIUM SYMBIOSIS
- PREDICTION OF EFFECTS OF SOIL WATER AND NITROGEN ON BARLEY QUALITY FOR MALTING USING A SIMULATION MODEL
- EFFECT OF LONG TERM APPLICATION OF PHOSPHORUS FERTILIZER ON BICARBONATE EXTRACTABLE SOIL PHOSPHORUS IN A MYWYBILLA SOIL
- CHANGES IN NITROGEN AND ORGANIC CARBON IN A MYWYBILLA CLAY FOLLOWING APPLICATIONS OF N FERTILISER RATES TO NINE CROPS
- EVALUATION OF FOUR ZINC FERTILISER SOURCES IN TWO SOILS FROM THE DARLING DOWNS, QUEENSLAND
- CAN VULPIA IN PASTURES BE CONTROLLED WITHOUT USING HERBICIDES?
- THE EFFECT OF GOAT MANURE AND SOIL MOISTURE CONTENT ON TILLER NUMBER AND LEAF YIELD OF BRACHIARIA DECUMBENS OVER THREE GROWTH CYCLES
- 648edmonson
- TECHNOLOGY TRANSFER OF EARLY SEEDING TECHNIQUESIN LOW RAINFALL CROPPING ENVIRONMENTS
- Development And Application Of Improved Selection Methods To Produce New Crops
- IMPROVEMENT PROGRAM FOR THE MILLET/PANICUM INDUSTRY
- CONTROL OF TAKE ALL OF WHEAT WITH CHLORIDE, BREAK CROPS, FUNGICIDE, FUMIGATION AND MICRO ORGANISMS
- THE EFFECTS OF A SULPHONYLUREA HERBICIDE ON ANNUAL MEDIC IN ALKALINE SOIL
- HOWWET: TRACKING FALLOW SOIL MOISTURE
- TOWARDS GENETIC MODIFICATION OF RUMEN BACTERIA FOR IMPROVED PASTURE FIBRE DIGESTION
- FORWARD ESTIMATES OF BARLEY PRODUCTION — A FEASIBILITY STUDY
- PASTURE STRATEGIES AND FERTILIZER TACTICS IN CEREAL PASTURE FARMING REWARDS AND RISKS
- ON FARM AND COMMUNITY MANAGEMENT STRATEGIES TO SUPPRESS MOUSE PLAGUES
- Productive Cropping Systems With High Water Use
- PASTURE CLEANING INCREASES CLOVER CONTENT OF SPRING PASTURE
- ECONOMIC EVALUATION OF NITROGEN FERTILISER ON OATS
- HIGH INPUT CROPPING: A VIABLE OPTION
- ESTIMATING SOIL WATER: TO KICK, TO STICK, TO CORE OR COMPUTE?
- TILLAGE AND STUBBLE EFFECTS ON WHEAT PRODUCTION IN TWO CONTRASTING SEASONS
- RELATIONSHIP BETWEEN YIELD OF GRAIN SORGHUM AND SOIL SALINITYUNDER FIELD CONDITIONS
- MODELLING NITRATE LEACHING UNDER SUGARCANE USING APSIM SWIM
- SWIMV2 IN APSIM: AN INTEGRATED PLANT, SOIL WATER AND SOLUTE MODELLING FRAMEWORK
- NATIONAL HERBICIDE RESISTANCE EXTENSION PROGRAM
- ANALYSING WHEAT BIOMASS AND GRAIN YIELD RESPONSE TO DROUGHT USING AFRCWHEAT2
- STYLOSANTHES SP. AFF. SCABRA, A NEW STYLO FOR HEAVY SOILS
- NITROGEN FIXATION BY POTENTIAL LEY PASTURE LEGUMES FOR CENTRAL QUEENSLAND: I GLASSHOUSE STUDIES
- NITROGEN FIXATION BY POTENTIAL LEY PASTURE LEGUMES FOR CENTRAL QUEENSLAND: II FIELD STUDIES
- AN EVALUATION OF ACACIA SALIGNA AS A MAINTENANCE FORAGEFOR SHEEP
- How Bad Is The Drought? Using A Cropping Systems Model To Assess Severity Of Drought
- How Bad Is The Drought? Using A Cropping Systems Model To Assess Severity Of Drought
- AN AUDIT OF FARMING AND FALLOW MANAGEMENT SYSTEMS IN SOUTH QUEENSLAND
- A MUNGBEAN CULTIVAR X POPULATION X ROW SPACING STUDY
- VARIATION IN NATURALISED MEDICAGO POLYMORPHA COLLECTED FROM INLAND EASTERN AUSTRALIA: 11 QUEENSLAND
- THE EFFECT OF WATERLOGGING ON THE PERFORMANCE OF PERENNIAL PASTURE
- TIMING OF DROUGHT DOES NOT AFFECT FIELD PEA YIELDS
- Catching Weed Seeds At Harvest: A Method To Reduce Annual Weed Populations
- MEASURING SPATIAL VARIATION WITHIN PASTURES
- SPATIAL ESTIMATION OF RAINFALL USING ARTIFICIAL NEURAL NETWORK
- ARE WHEAT SEEDLINGS INTELLIGENT?
- 689Miyan
- LIGHT INTERCEPTION AS A FACTOR IN SEED AND GUM YIELD OF GUAR
- HERBICIDE RESISTANCE IN WILD OATS AND ANNUAL RYEGRASS
- Crop Rotation Effects On A Soil In Summer Rainfall Australia
- WINDBREAK RESEARCH IN SOUTH AUSTRALIA
- SOIL EVAPORATION ESTIMATION: A COMPARISON OF APPROACHES
- AGRONOMIC FACTORS AFFECTING PROTEIN CONTENT OF IRRIGATED SOYBEANS IN NORTHERN VICTORIA
- PERENNIAL RYEGRASS AND TALL FESCUE RESPONSES TO WATER DEFICITS
- LIMITATIONS TO NITROGEN FIXATION IN WHITE CLOVER DAIRY PASTURES
- CONSERVATION TILLAGE PROVIDES BONUS CROPS
- VERTICAL INTEGRATION FROM PADDOCK TO PLATE: ADZUKI BEANS
- VERTICAL INTEGRATION FROM PADDOCK TO PLATE: ADZUKI BEANS
- GROWTH RESPONSES OF THREE SUMMER LEGUME SPECIES
- Multimedia Campaign For Agricultural Extension
- COMPOUND TRACE ELEMENT FERTILISERS SUPERIOR TO DRY BLENDS
- SHEEP RESPONSE TO SUPER PHOS
- THE EFFECT OF NITROGEN AND DEFOLIATION TIMING ON OAT / MEDIC MIXTURES
- PRISM: APPLYING BIOECONOMIC MODELS TO FARM MANAGEMENT ISSUES
- EFFECT OF NURSERY FERTILIZER APPLICATION AND SEEDLING AGE ON THE GROWTH AND YIELD OF A TRADITIONAL RICE CULTIVAR IN THE RAINFED LOWLANDS OF CAMBODIA
- COMPARATIVE WATER USE OF CROP ROTATIONS ON A SANDPLAIN SOIL.
- THE EFFECT OF PROMOTING EARLY VIGOUR ON WHEAT YIELDS UNDER THREE ENVIRONMENTAL REGIMES
- SPREADING THE WORD IN AGRONOMY
- CROP MONITORING EVALUATING FARMING PRACTICES IN A MARGINAL CROPPING ZONE
- TARGETING CLIMATE FORECASTS TO CROPPING IN NE AUSTRALIA
- THE BENEFITS OF TREES FOR IMPROVING THE VALUE OF GRAZING PASTURES IN THE UPPER SOUTH EAST OF SOUTH AUSTRALIA
- MATCHING NITROGEN MANAGEMENT TO SOILS AND CLIMATE A STRATEGIC ROLE FOR MODELS
- EFFECT OF PREVIOUS CROP ON IRRIGATED WHEAT YIELD
- ARE NITROGEN RESPONSES OF SPACED WHEAT PLANTS REPRESENTATIVE OF A FIELD CROP?
- Tall Fescue Breeding And Improvement
- WHITE CLOVER IMPROVEMENT AND BREEDING
- MODELLING NITROGEN LOSSES UNDER SUGARCANE CROPS USING APSIM SWIM
- WATER BALANCE AND DRAINAGE IN DUPLEX SOILS
- VEGETABLE GROWING SYSTEMS: IMPACTS ON THE ENVIRONMENT
- THE POTENTIAL TO USE CARBON ISOTOPE DISCRIMINATION AS A SELECTION TOOL TO IMPROVE WATER USE EFFICIENCY IN SOYBEAN
- MODEL ANALYSIS OF VARIABILITY OF MAIZE POTENTIAL PRODUCTIVITY IN A COOL CLIMATE
- EFFECTS OF METHANOL ON GROWTH, WATER USE AND YIELD OF BARLEY
- THE ROUGH AND SMOOTH OF RAINFALL CAPTURE IN THE MARANOA
- THE APPLICATION OF TOTAL QUALITY MANAGEMENT PRINCIPLESTO SUGARCANE RESEARCH AND EXTENSION PROGRAMS
- COMPETITION AND THE ESTABLISHMENT OF ALEMAN GRASS
- SELECTION FOR WATER USE EFFICIENCY IN GRAIN LEGUME SPECIES
- Contributed Papers
- 1998 9th AAC
- Pasture Species
- Effects Of Pasture Management On Germinable Seed Bank In A Degraded Phalaris Pasture 1
- A Bromoxynil Tolerant Transgenic Subterranean Clover 1
- Persistence Of Late Maturing Subterranean Clover Cultivars In Western Australia 1
- A Comparative Performance Of Early Flowering Subterranean Clover Genotypes 1
- Subtropical Grass Evaluation For Pastures In Northern NSW 1
- Imbibition And Germination Of Yellow Serradella Seeds 1
- Effect Of Temperature And Light On Seed Softening In Yellow Serradella 1
- Evaluation Of Temperate Perennial Grasses For Low To Medium Rainfall Areas Of Tasmania 1
- Evaluation Of Trifolium Subterraneum Ssp. Brachycalycinum In South Australia, New South Wales And Queensland. 1
- Biodiversity And Potential Utilisation Of Blown Grasses (Agrostis Spp.) In Lowland Victoria 1
- Performance Of Pasture Legumes On Three Contrasting Soil Types In Western Victoria 1
- Breeding Perennial Lablab: Performance Of Selected Lines In Queensland 1
- Anthracnose Resistance In Stylosanthes Seabrana 1
- Yield Of Lines Bred To Improve Anthracnose Resistance In Stylosanthes Scabra 1
- Early Flowering Balansa Clovers For The Cereal Livestock Zone 1
- Seedling Establishment, Plant Survival And Response To Grazing Of New Acid Soil Tolerant Phalaris Populations 1
- Carbon Isotope Discrimination And Specific Leaf Weight Estimate Transpiration Efficiency Indirectly In Stylosanthes Under Well Watered Conditions 1
- The Evaluation Of Tropical Legumes For Use In Ley Pastures In Central And Southern Queensland 1
- Subterranean Clover Seed Bank Responses To Pasture Management 1
- Fate Of Serradella, Medic And Biserrula Seeds In Pods Ingested By Sheep 1
- Genetic Variation In Trifolium: Comparison Of Some Outcrossing And Inbreeding Species From Turkey 1
- Above And Below Ground: A Comparative Study Of The Root And Shoot Growth Of Pasture Legumes 1
- Evaluation Of Annual Pasture Legume Species For Acidic, Deep Sandy Soil. 1
- Contributed Papers
- Poster Papers
- Pasture Management
- Measurement Of Evapotranspiration From Different Pasture Types Using The Rapid Chamber Method 1
- Breeding Better Lucernes For The Cropping Zone Of Eastern Australia 1
- Effects Of Sub Optimal Irrigation Practices On Dairy Pasture Production In South West Victoria 1
- A Strategic Assessment Of Sustainability Of Grazed Pasture Systems In Terms Of Their Water Balance 1
- Established Phalaris And Cocksfoot Reduce The Growth, Turgor And Survival Of Subterranean Clover Seedlings 1
- The Red Barren Project: When Salt Is Not Salt 1
- Comparative Pasture Growth And Soil Profile Dryness Of A Lucerne And An Annual Legume Pasture 1
- Modelling Lucerne Growth Using APSIM 1
- The Effect Of Pasture Type And Management On Soil Matric Potential And Root Profile 1
- Salinity Effects On Irrigated Lucerne 1
- Water Usage And Dry Matter Production Of Perennial Pasture Species Down A Duplex Toposequence 1
- Contributed Papers
- Poster Papers
- Pasture Management
- Using The Grassgro™ Decision Support Tool For Grazing Systems 1
- Response To Superphosphate By Pasture Grazed By Sheep In A Cell System 1
- Pasture Ready Reckoner: Group Learning For Woolgrowers Through Benchmarking 1
- Using GrassGroTM; To Support Tactical Decisions On Grazing Farms: A Case Study At "Yaloak", Ballan, Victoria 1
- Botanical Composition Responses To Grazing Pasture By Different Classes Of Sheep 1
- Native Pasture Management In The 400 600 Mm Rainfall Zone Of Northern NSW 1
- The Implementation And Limitations Of Using A Falling Plate Meter To Estimate Pasture Yield 1
- Tolerance Of Tree Seedlings To Pre And Post Emergence Herbicides 1
- Emergence And Establishment Of Wallaby Grass (Danthonia Spp.) 1
- Oversowing Legumes Into Perennial Pasture: Can We Do It Better? 1
- Biodiversity, Stability And Pasture Management The Role Of Functional Groups 1
- Comparison Of Spring Sown Brassica Spp. In Direct Drilled And Cultivated Seedbeds 1
- Farmer Adoption Of Dryland Lucerne Systems In North Central Victoria 1
- Sustainable Grazing Systems (SGS) – Developing A National Experiment 1
- Establishment Of Undersown Lucerne In Southern NSW 1
- Effects Of Grazing Management On Productivity Of Perennial Ryegrass Pastures 1
- Nutrition And Disease Control Improve Victorian Mallee Pastures 1
- 106kemp
- Pasture Management For Both Production And Stability 1
- Aboveground And Belowground Competition Among Pasture Species 1
- Effect Of Defoliation Management On Component Dry Matter Yield Of Kikuyu(Pennisetum Clandestinum) Grass 1
- Factors Affecting The Incidence Of The PE "sudden Death" Form Of Phalaris Aquatica Poisoning In Sheep. 1
- Effect Of Nitrogen On Pasture Yield And Quality For Silage In Western Victoria 1
- Contributed Papers
- Poster Papers
- Summer Crops
- Efficient Screening For Mid Season Cold Damage In Rice 1
- Varietal Response To Mid Season Cold Damage In Australian Rice 1
- Variation In Radiation Use Efficiency In Temperate Rice 1
- Impact Of Agronomic Practices On Cadmium Uptake By Peanuts 1
- Genotypic Variation For Rate And Onset Of Leaf Senescence In Grain Sorghum 1
- Premature Senescence Of Cotton The Future Of Crop Nutrition Problems? 1
- Use Of Thermal Parameters To Reduce The Number Of Sugarcane Clones Required For Trash Blanketing Trials 1
- Early Silage Maize For Cool Summers? 1
- Genotypic Variation For Grain Yield Of Rice Under Water Deficit Conditions 1
- A New Sorghum Planting Strategy Resulting From The Synergy Of Farmer Systemic Knowledge And Systematic Simulation Of Sorghum Production Systems 1
- Leaf Nitrogen Gradients In Cotton Canopies Vary With Ontogeny And Nitrogen Supply 1
- Growth Analysis Of Short And Long Season Cotton Cultivars? 1
- Physiological Studies Of Ratooned And Non Ratooned Winged Bean Grown With And Without Support 1
- Alleviation Of Drought Stress And Aflatoxin Incidence In Peanut Using Short Maturing Varieties 1
- A Germplasm And Breeding Strategy For Improved Drought Resistance In Phaseolus Vulgaris 1
- Predicting Mungbean Response To Water Stress 1
- Allelopathic Behaviour Of Grain Legumes In Cotton Based Farming Systems 1
- Improving Drought Tolerance Of Cotton By Glycinebetaine Application And Selection 1
- Fibre Development In Early Harvest Ginger Grown In South East Queensland 1
- Lima Beans Potential Grain Crop For Southern Queensland 1
- Varietal Characteristics For Adaptation Of Cotton (Gossypium Hirsutum L.) To Raingrown Conditions 1
- Yield Performance Of Seven Rice Populations In Rainfed Lowland Systems In Thailand 1
- Spikelet Sterility Induced By Nitrogen Fertilisation And Low Temperature In Rice 1
- Yield And N2 Fixation Of Backcrossed Supernodulating Soybean Mutants
- Osmotic Adjustment Of Sorghum Genotypes Under Two Nitrogen Levels In The Field 1
- The Growth And Yield Of Cotton Following Seven Different Rotations 1
- Predicting Sweet Corn Maturity For Factory Processing In Canterbury, New Zealand 1
- Drought Effects On Canopy Development In Sweet Corn 1
- Towards A Robust Method Of Modelling Leaf Appearance In Plants 1
- Drought Effects On Water Use, Growth And Yield Of Sweet Corn 1
- Row Spacing Effects On Two Cultivars Of Mungbean (Vigna Radiata) At Gatton 1
- Effect Of Nitrogen And Cutting On Forage And Grain Characters Of Millets 1
- Contributed Papers
- Poster Papers
- Winter Crops
- Developing Faba Beans As A Viable Industry In Southern New South Wales 1
- Low Adoption Of Lupins In Southern NSW Insights From PRISM 1
- Towards Better Canola Yield: A Principal Components Analysis Approach 1
- Canopy Architecture, Light Utilisation And Productivity Of Intercrops Of Field Pea And Canola 1
- Productivity And Water Use Of Intercrops Of Field Pea And Canola 1
- Variation In Yield Of Commercial Field Pea Crops In Southern NSW 1
- Pulses Increase Profitability Of Winter Crop Rotations In Southern NSW 1
- Drought Stressed Mustard Yields More Than Canola Due To Greater Leaf Turgor 1
- Early Generation Yield Performance Of Different Field Pea Plant Types 1
- ShowDevel: Predicting And Visualising Wheat Development For Planning And Management 1
- Cultivar Variability In Acidic (Al/Mn) Stress Tolerance In Triticale 1
- Exudation Of Organic Acids From Roots Of Triticale 1
- Water Use Efficiency Of Canola In Victoria 1
- Effects Of Early Water Deficit On Growth And Development Of Faba Bean 1
- Management Of Winter Wheat For High Forage And Grain Yields 1
- Effect Of Plant Density, Sowing Date And Irrigation On The Yield Of Fibre Hemp (Cannabis Sativa) And Flax (Linum Usitatissimum). 1
- The Potential For Using Hemp (Cannabis Sativa) And Flax (Linum Usitatissimum) Fibre As A Reinforcing Agent In Newsprint Manufacture. 1
- Grain Growth Within Oat Panicles 1
- Implications Of Accounting For Below Ground N On The Calculations Of Residual Returns Of Fixed N For Commercial Faba Bean Crops 1
- Evaluation Of A Germplasm Collection Of Fenugreek (Trigonella Foenum Graecum). 1
- 228martin
- 229martin
- Coordination Of Apex And Leaf Development In Oats 1
- Assisting New White Lupin Processing Industries By Breeding Larger And Smaller Seeded Cultivars 1
- Field Pea Yields Are Declining In Medium To High Yielding Areas 1
- A Breeding Solution To Improving Seedling Establishment Of Wheat 1
- Determination Of Grain Protein Concentration In Barley 1
- Evaluation Of Legume And Cereal Fodder Species For Variation In Water Use. 1
- Opportunities And Threats To Temperate Oilseed Production In Australian And Canadian Farming Systems 1
- Increased Market Share For Australian Pulses: A Quality Assurance Strategy 1
- Biofumigation By Brassicas Reduces Take All Infection 1
- Agronomy Of Lathyrus Species In South Australia 1
- Soil Biological Factors Limit The Early Growth Of Direct Drilled Wheat 1
- Effect Of Harvest Date On Quality And Dry Matter Of Cereal Cultivars 1
- Effect Of Tillage On Rhizosphere Microbial Populations, Early Growth And Grain Yield Of Wheat Throughout South Eastern NSW 1
- Effects Of Genotype And Environment On Grain Filling In Barley Grown In South East Australia 1
- Contributed Papers
- Poster Papers
- Weed Control
- Changes In Pasture Management Needed For Improved Control Of Weeds In Pastures 1
- Can Cattle Spread Giant Rats Tail Grass Seed (Sporobolus Pyramidalis) In Their Faeces? 1
- Changes In Pasture Management Needed For Improved Control Of Weeds In Pastures 1
- Winter Cleaning Subterranean Clover Pastures With Non Selective Herbicides 1
- Management Of Wild Oats And Paradoxa Grass With Reduced Dependence On Herbicides 1
- Pasture Plant Back Periods Following Application Of Simazine To Control Vulpia Spp. 1
- Tordon* Herbicides Old Answers For New Problems 1
- Herbicide Resistance In Wild Oats In Southern New South Wales 1
- Differential Allelopathic Potential Among Wheat Accessions To Annual Ryegrass 1
- Can Fire Control Giant Rats Tail Grass (Sporobolus Pyramidalis)? 1
- Persistence Of Sulfonylurea Herbicides On Alkaline Soils. 1
- The Ecology Of Wild Radish : The Key To Sustainable Management 1
- Competition Between Rampion Mignonette (Reseda Phyteuma L.) And Wheat, Beans, Fescue And Subterranean Clover 1
- Herbicide Control Of Mintweed (Salvia Reflexa) In Cotton Production Systems 1
- Evaluating The Accuracy Of Weed Maps Generated Using Airborne Multispectral Imaging 1
- Herbicide Resistance In Wild Radish (Raphanus Raphanistrum): New Seed Soaking Method For Resistance Testing 1
- Allelopathy: From Concept To Reality 1
- Comparison Of Three Weed Control Methods: Chemical, Flame And Hot Water. 1
- Contributed Papers
- Poster Papers
- Farming Systems
- Crops And Pastures A Profitable Combination For The Central Tablelands Of NSW? 1
- Managing Annual Pastures To Benefit Grain And Out Of Season Lamb Production 1
- The Future Of Agriculture In The Peri Urban Fringe Of Sydney 1
- Changes In Botanical Composition Under Free Range Chicken Farming 1
- Permacropping Into A Perennial Native Pasture Base 1
- Sustainable Agriculture Using Best Practice To Manage The Paradoxes Facing Land Managers And Agronomists 1
- A Framework For Extending Principles Of Conservation Cropping 1
- Nitrogen Fertiliser Decisions For Wheat On The Liverpool Plains, NSW. II. Should Farmers Consider Stored Soil Water And Climate Forecasts ? 1
- The Use Of Seasonal Climate Forecasts In Cropping Systems Management 1
- Variations In Wheat Yields Behind Windbreaks In Southern Queensland 1
- Extension And Demonstration The Final Keys For Successful Adoption Of Conservation Tillage 1
- Evaluation Of Wagga Wagga Shire Wheat Yields Using Simulated Production And Water Balance 1
- Management Of Prime Quality High Protein Wheat In South Australia 1
- Comparisons Of Farming Systems Yield More Questions Than Answers 1
- Organic Farming Now And The Future 1
- Gravel And Conventional Mole Drainage For Dryland Cropping In SE Australia 1
- FARMSCAPE Online: Using The Internet For Supporting Interactions Among Farmers, Advisers And Researchers 1
- Evaluation Of Participative Approaches To RD&E: A Case Study Of FARMSCAPE 1
- The FARMSCAPE Approach To Farming Systems Research 1
- 199liu
- Increasing The Benefits Of Irrigation On Cropping Land In The Southern Murray Darling Basin 1
- Community Expectations And Perceptions Of Agriculture In Peri Urban Regions 1
- Shelterbelts And Wheat Production In North East Victoria 1
- Nutritional Status Of Wheat On Acid And Limed Soils In Southern NSW 1
- A Farming Systems Study At Nindigully, South West Queensland 1
- The Effect Of Nitrogen And Phosphorus Fertilisers On Barley In Southern Australia 1
- Wheat Yield Trends In The Northern Grains Region A Sustainable Industry? 1
- The Management Of Models: Risks And Responsibility 1
- Farming Styles And Extension In Broadacre Cropping 1
- Airborne Multispectral Imaging And Precision Farming The South Eastern Australian Experience 1
- Contributed Papers
- Poster Papers
- Soil Fertility
- Describing Soil PH Profiles And Their Effect On Yield Of Cereal Crops 1
- Development Of A Carbon Based Sustainability Index For Ferrosols In Rainfed Cropping Systems 1
- Phosphorus And Carbon Losses Off Dairy Catchments Located On Duplex Soils 1
- Changes In Soil Ph Resulting From Simulated Urine Patches 1
- Nutrient Export From Rural Land In The Hawkesbury Nepean Catchment 1
- Burial Of Lime By Earthworms In Acid Soils In Western Australia 1
- Soil Acidification Prediction And Quantification Using APSIM SWIM. 1
- Minimum Tillage Effects On Soil Structure Measured Using Image Analysis 1
- Evolution Of Nitrate And Ammonium From Banded Urea At Two Soil Temperatures 1
- Impact Of Cropping On Fertility Of North Queensland Ferrosols
- The Impact Of Rotation, Stubble Management And Tillage On The Relative Contribution Of Bacterial And Fungal Microbial Biomass. 1
- Nitrogen Fertiliser Decisions For Wheat On The Liverpool Plains NSW 1. Should Farmers Consider Paddock History And Soil Tests? 1
- Effect Of Subsoil Acidity Treatments On The Chemical Properties Of A Ferrosol 1
- Soil Physical Status In A Long Term Land Management Trial And Adjacent New Trial At Halbury, South Australia 1
- Factors Influencing Soil And Nutrient Loss In Stormwater From A Market Garden 1
- SOILpH A New APSIM Module For Management Of Soil Acidification 1
- Improving Management Using Soil Monitoring And Simulation: A Case Study In Co Operative Learning 1
- Characterising Soils For Plant Available Water Capacity Challenges On Shrink/swell Soils 1
- Piggery Bedding Litter As A Source Of N For Field Crops 1
- Soil Health And Crop Productivity In Alfisols With Integrated Plant Nutrient Supply System 1
- Nitrate Leaching Under Sugarcane: Interactions Between Crop Yield, Soil Type And Management Strategies 1
- Soil PH Change Over Time In Relation To Rotation, N Fertiliser, Stubble Management And Tillage 1
- 277michalk
- Effect Of Various Management Systems On N Mineralisation In Surface Soils 1
- N Mineralisation And Nitrification In Crop And Pasture Soils 1
- Dairy Pasture Yield And Quality Responses To Nitrogen, Phosphorus, Potassium And Sulphur 1
- Contributed Papers
- Poster Papers
- Rotation Systems
- Response Of Wheat Yield To Preceding Crop And Fertiliser Nitrogen 1
- Effects Of Different Pastures On Soil Quality 1
- A Survey Of Methods Of Lucerne Removal Prior To Cropping 1
- Role Of A Grass/medic Ley In The Cropping System Of North Western NSW 1
- Using Ley Pastures To Improve Infiltration 1
- Comparisons Of The Efficiency Of Nitrogen Fixation In Pastures 1
- High Yielding Crops From Legume Dominant Pastures 1
- Long Term Residual Effects Of 6 Different Crops On Yield And Protein Of Wheat 1
- Management Options For Pasture And Cereal Production In North West Victoria 1
- Hydrologic Utility Of Phase Farming Based On Winter Rainfall In South Eastern Australia. 1
- Pasture Species And Phase Length Effects On Rice Grain Yield. 1
- DACS: A Knowledge Based Decision Support System For Dryland Agriculture Crop Sequencing 1
- Managing Residues And Fertilisers To Enhance The Sustainability Of Wheat Cropping Systems 1
- Simulating Peanut/Wheat Cropping In The Burnett With APSIM 1
- Stubble Management Influences Nitrogen Uptake And Yield In Subterranean Clover/wheat Rotations 1
- Phase Farming Using Lucerne In Crop Rotations 1
- The Timing Of Lucerne Removal In The Transition To A Cropping Phase 1
- Participatory Research Using On Farm Monitoring And Simulation: Spring Sown Mungbeans 1
- Lucerne In Crop Rotations 1
- Predicted Consequences Of Early Application Of Nitrogen Fertiliser 1
- Profitable Double Cropping Rotations Involving Cereals And Pulses In Central Queensland 1
- Crop Rotations Increase Productivity In No Tillage Systems In Northern New South Wales
- Fertiliser And Legume Residues Effect On The Productivity Of Irrigated Rice Wheat Systems In Bangladesh 1
- Contributed Papers
- Poster Papers
- Conference Information
- Plenary Papers
- The Donald Oration
- The Environmental Imperative
- The Importance Of High Farm Yields To Wildlife Conservation
- Biotechnology Risks And Benefits: The Ingard Cotton Example
- Farming Fragile Environments: Low Rainfall And Difficult Soils In South Australia
- Looking For Opportunity And Avoiding Obvious Potholes: Some Future Influences On Agriculture To 2050
- The State Of The Australian Environment
- Managing Soil Water And Nitrogen To Minimise Land Degradation
- The Future Challenge
- The Quality Imperative
- The Socio Economic Imperative
- Women: The Silent Partners Of Agriculture
- Responding To The Wool Crisis With A Focus On Technology Adoption
- Technology Transfer Perceptions Of Extension Agronomists
- Learning From Farmers The Check Approach
- Who Controls Our Future?
- Agriculture And The Economy
- Land Degradation Issues And Management Concerns For Aboriginal Communites Of Central Australia
- Pasture Species
- 2001 10th AAC
- Concurrent Session 1, 1300 1430, Monday, 29/1/2001
- Precision Farming, Controlled Traffic, Spatial Variability
- Mechanising On The Run Grain Sampling 1
- Strategies To Interpret The Yield Map: Defining Yield Limiting Factors 1
- Yield And Protein Variation Within A Controlled Traffic System 1
- Strategies To Interpret Yield Maps: Predicting Grain Protein Using Yield 1
- Understanding Subsoil Water Use On Southern Mallee Soils: I. Spatial Characteristics Of Subsoil Constraints 1
- Understanding Subsoil Water Use By Cereals On Southern Mallee Soils: II. Crop Response 1
- Age Of Cultivation Is Not A Good Indicator Of Profitable Crop Production 1
- Innovative Management Of Grain Sorghum In Central Queensland. 1
- Drought And Water Use Efficiency
- Genotypic Variation In Water Potential At Different Positions And Water Conductance In Rice 1
- Effect Of Timing And Intensity Of Drought On The Yield Of Oats (Avena Sativa L.) 1
- Defoliation Of Sorghum And Wheat Leads To Control Of Phenology And Enhanced Water Use Efficiency. 1
- Terminal Drought And Seed Yield Of Lupin 1
- Management Practices To Reduce Aflatoxin Contamination In Peanut 1
- Relations Among Yield Potential, Drought Tolerance And Stability Of Yield In Bread Wheat Varieties Under Water Deficit Conditions 1
- Influence Of Cultivar Maturity And Leaf Type On The Agronomic Water Use Efficiency Of Raingrown Cotton 1
- Adaptation Of Chickpea To Water Limited Environments 1
- Differential Responses To Water Regimes In Triticale Genotypes Differing In Aluminium Tolerance 1
- Nutrition: Nitrogen Management
- The Nitrate Scavenging Ability Of Phalaris And Lucerne In Subterranean Swards 1
- Contribution Of Pasture Legumes To Soil Mineral Nitrogen In Northern Victoria 1
- A Monitoring System Based On Amino N At Harvest Time To Improve Nitrogen Management In Sugarcane Systems 1
- Nitrogen Supply And The Establishment And Productivity Of Chicory (Cichcrium Intybus L.) Based Pastures 1
- Comparison Of KCl 40 And MCP S In Predicting S Response In Grain SorghumGrain Sorghum A Yield Response To N Or S? 1
- Nitrogen Response Efficiencies From Grazed Dairy Pastures Under Dry Soil Conditions 1
- Contributions Of Fixed Nitrogen By Crop Legumes To Farming Systems Of Eastern Australia 1
- Forage Production And Nitrogen Uptake Of Forage Sorghum, Grain Sorghum And Maize As Affected By Cutting Under Different Nitrogen Levels 1
- Impact Of Nitrogen Management And Subsoil Properties On Deep Nitrate Accumulations In Soils Of The Darling Downs: Case Studies 1
- Phenology And Crop Development
- Effect Of Temperature On The Rate Of Early Fruiting Developmental Processes Of Cotton 1
- The Cost Of Low Temperature To The NSW Rice Industry 1
- Sowing Dates, Phenology And Yield In Lupins (Lupinus Angustifolius) 1
- Growth, Dry Matter Production And Seed Yield Of Indian Mustard (Brassica Juncea L.) In The Mediterranean Environment Of South Western Australia 1
- The Effect Of Planting And Harvest Time On Sugarcane Productivity 1
- Title: Assessing The Risk Of Cotton ‘earliness’ Management Strategies With Crop Simulation. 1
- Phenology Of Canola Cultivars In The Northern Region And Implications For Frost Risk 1
- Evaluation Of Simtag And Nwheat In Simulating Wheat Phenology In Southeastern Australia 1
- Precision Farming, Controlled Traffic, Spatial Variability
- Concurrent Session 2, 1600 1730, Monday 29 January
- Farming Systems 1: Rotations, N And Disease
- Yield Responses To Breaking The Sugarcane Monoculture 1
- Effect Of Soil Borne Ascochyta Blight Fungi On The Grain Yield Of Field Peas 1
- Growth Of Different Plant Species In Pasteurised/fumigated And Untreated Sugarcane Soils 1
- Species And Management Of Fallow Legumes In Sugarcane Farming Systems 1
- Short Term Rotations Using The Forage Legume Lablab Have A Place In Central Queensland Farming Systems 1
- Survey Of Field Pea Production Practices In South Australia. 1
- Blackspot Survival In Soil And Stubble And Aerial Dissemination Through The Season 1
- Agronomic Management For Production Of Prime Hard Quality Wheat In South Australia. 1
- The Effect Of Brassica Crops On The Level Of Mycorrhizal Inoculum In Soil 1
- Water Management
- What Is Limiting Productivity And Water Use Of Cereals In The Southern Wimmera Of Victoria? 1
- Raised Bed Farming Of Waterlogged Duplex Soils In Western Australia 1
- Soil Properties Under Raised Bed Farming Systems In Tasmania 1
- The Adoption Of Sub Surface Drainage And On Off Grazing By Victorian Dairy Farmers 1
- Soil Water Balance Modelling Highlights Limitations For Pasture Production In Northern NSW 1
- Soil Water Characteristics Of A Red Chromosol And Brown Vertosol And Pasture Growth 1
- Plant Density, Litter And Bare Soil Effects On Actual Evaporation And Transpiration In Autumn 1
- Long Term Effects Of Pugging On Soil And Pasture In SW Victoria 1
- Agronomic Responses Of Cotton To Low Soil Oxygen During Waterlogging 1
- Nutrition: P And Micronutrients
- Management Effects On Cadmium Accumulation By Peanut And Soybean 1
- Narbon Bean Response To Fertiliser Nutrients In The Victorian Mallee 1
- Effect Of Superphosphate On Production And Botanical Composition Of Native Grass Pastures In Tasmania 1
- Effects Of Zinc On Photosynthesis And Yield Of Wheat Under Heat Stress 1
- Using Phosphorus Budgets To Assess Fertiliser Use In Vegetable Cropping In The Sydney Basin 1
- Agronomic Effectiveness Of Selected Fertilisers In Supplying Copper To Perennial Ryegrass 1
- Soil Fertility And Wheat Crop Response To Phosphorus Fertiliser On Vertosols In Low Rainfall Areas Of The Northern Grain Zone. 1
- Effects Of Phosphate Fertiliser And Perennial Pastures On Sheep Production In Northern Victoria 1
- The Effect Of Hybrids, Soil Types And Applied Phosphorus On The Growth And Tissue Composition Of Pyrethrum (Tanacetum Cinerariifolium L) 1
- Crop Physiology And Management
- Performance Of Modern Wheat Cultivars As Affected By Leaf Area, Disease And Weather Conditions 1
- Effects Of Sowing Time On Pyrethrins Yield Of Pyrethrum (Tanacetum Cinerariifolium) In Tasmania 1
- Physiological And Morphological Responses Of Tall Fescue And Perennial Ryegrass To Leaf Defoliation 1
- Potential Alternative Oilseeds For South Eastern Australia 1
- Fruit Production Rates In Cotton Cultivars Of Different Maturity Times 1
- Physiological Basis Of Grain Size Variation Amongst Barley Cultivars 1
- Changes In Sugar Content Of Australian Sugarcane Cultivars After Application Of Chemical Ripeners 1
- The Importance Of Second Flushes Of Flowers In Mungbean Production 1
- Farming Systems 1: Rotations, N And Disease
- Concurrent Session 3, 1130 1300, Tuesday 30 January
- Weather Forecasting, Environmental Risk Management
- Improving The Reliability Of Sorghum Production In The Farming System 1
- Production Of Legume Forage Crops In Frost Prone Areas Of Tasmania 1
- Nitrogen Decreases Deep Irrigation Efficacy In Reducing Low Temperature Damage In Rice 1
- Temperature Variation And Frost Risk In Undulating Cropland 1
- Urban Encroachment And Loss Of Prime Agricultural Land 1
- From Oceans To Farms – The Agricultural Potential Of Ocean Based Forecasts 1
- The Influence Of Climate Variability On Cropping Systems In Central Queensland 1
- An Alternate Method For Estimating The Value Of The Southern Oscillation Index (SOI) In The Northern Grainbelt 1
- Risk Assessment And Expert Opinion In Implementing Policy Exceptional Circumstances. 1
- Managing Soil Problems: Salinity, Acidity
- Soil Structure Affects Water Balance Of Ferrosol Cropping Systems 1
- Pasture Responses To Lime Over Five Years Are Limited And Highly Variable 1
- Melilotus Alba: The Preferred Forage Legume For Autumn And Spring Summer Production On Saline Soils In SW Victoria. 1
- Lucerne Varieties Differ In Their Response To Liming On An Acid Soil. 1
- Salinity And Sodicity, Implications For Farmers In Central Queensland 1
- The Effects Of Changed Subsoil Structure On Yield And Yield Quality 1
- A Case Study To Reduce Dryland Salinity On A Temora Farm 1
- Impacts Of Surface Applied Lime On Sheep Production Systems In South Western Victoria 1
- Soil Acidity In The High Rainfall Wheat Belt Of Southern NSW. 1
- The Effect Of Various Calcium Amendments Applied To Sugarcane Trash 1
- Decision Support Systems And Fertiliser Response
- Phosphorus Fertiliser For Beef Or Sheep Pastures A Decision Support Package 1
- Modelling Strategic Nitrogen Fertiliser Decisions For Central Queensland Farming Systems 1
- Testing NutriLOGIC, A Decision Aid For Nitrogen Fertiliser Management In Cotton 1
- Comparison Of Single High Rate And Multiple Low Rate Zinc Fertiliser Applications To Increase Available Soil Zinc Supplies. 1
- Comparing The Effects Of TSP, MAP And DAP Applied In The Seed Furrow On Establishment Of 5 Summer Crop Species. 1
- Comparing Effects Of TSP, MAP And DAP Applied In Bands On Some Key Soil Chemical Properties In A Black Vertosol. 1
- Fertiliser Adviser Crops: An Expert System For Tasmanian Crops 1
- Fertiliser Adviser Dairy An Expert System For Dairy Pastures In Tasmania 1
- Cumulative Effect Of The Application Of N And P Fertilizers On Soil Total And Labile Concentrations After 12 Cereal Crops On A Black Vertosol. 1
- Mid Infrared Spectroscopy For Rapid And Cheap Analysis Of Soils 1
- Pasture Cultivars And Management
- Stolon Formation In Perennial Ryegrass May Aid Persistence 1
- Trigonella Balansae – A New Pasture Legume For The Alkaline Soils Of Southern Australia? 1
- A Comparison Of The Emergence Of Octoploid And Tetraploid Phalaris Aquatica Seedlings 1
- Reducing The Cost Of Lucerne Establishment In Western Australia 1
- Managing Sirosa Phalaris In A Predominantly Summer Rainfall Environment 1
- A Field Kit For Producers To Assess Pasture Health In The Paddock 1
- Effects Of Pasture Type And Management On The Clover Content And Nutritive Value Of Dairy Pastures In South West Victoria 1
- A Monitoring Protocol For Annual Ley Pastures 1
- Frontier Balansa Clover Offers Exciting Prospects For Low Rainfall Pastures In Western Australia 1
- Weather Forecasting, Environmental Risk Management
- Concurrent Session 4, 1050 1220, Wednesday 31 January
- Farming Systems 2: Rotations, Legumes, Other Crops
- Effect Of Gypsum And Stubble Retention On Crop Productivity In Western Victoria 1
- Ley Cropping Systems Benefit From Low Rates Of Seed Softening Of Annual Pasture Legumes In Southern Australia. 1
- Assessing The Potential Of Wide Row Farming Systems In The Mallee 1
- Productive Lucerne Crop Rotations 1
- Better Pasture, Better Crop, Better Systems 1
- Economic Potential Of Ley Pasture Legumes For Central Western NSW 1
- Fenugreek (Trigonella Foenum Graecum) Compared To Five Temperate Legume Species In Wimmera Farming Systems. 1
- Effects Of Lucerne And Medics On Subsequent Crops In The Victorian Mallee 1
- Participatory Research, Action Learning
- Working With Farmers To Benchmark High Yielding Durum Wheat On The Liverpool Plains 1
- Experiences From Using Action Learning Groups To Develop Sustainable Farming Systems For Central Queensland 1
- Challenges Of On Farm Research – Insights From The Central Queensland Farming Systems Experience 1
- Impact Of An Action Learning Activity On Improved Nitrogen Management In Southern Queensland. 1
- Taking The Reigns: Farmer Driven Learning, Innovation And Extension For Profitable Dairy Pasture Production In New South Wales 1
- Success Factors For Participatory Farming Systems Projects – Field Notes From The North 1
- Paddock Selection Is Critical For Reliable Malt Barley Production 1
- Collaborating With Farmers To Design A Practical Soil Health Checklist 1
- A View Of Science, Agricultural Science And Farming Systems Research In These Postmodern Times 1
- Farming Systems 3: Science And Crop Monitoring
- Wheat Stubble Management Influences Emergence And Growth Of Canola 1
- Women ‘TopCroppers’ ~ A Valuable Part Of The Farm Workforce 1
- The Impact Of A Farming Systems Approach On Adoption Of Butterfly Pea In Central Queensland 1
- TOPCROP State Focus In Victoria. An Extension Program Tackling Industry Issues 1
- The Development Of Ecological Performance Indicators For Sustainable Systems 1
- Poor Wheat Yield Response To Conservation Cropping – Causes And Consequences During 10 Years Of The Harden Tillage Trial 1
- Agronomic Interactions Between Drought And Crop Sequence 1
- Warm Season Cropping In The Southern Cropping Zone Of Australia 1
- Cereal Yields Associated With Changes In Soil Characteristics Following Six Years Of Acacia 1
- Farming Systems 2: Rotations, Legumes, Other Crops
- Concurrent Session 5, 1600 1700, Wednesday, 31 January
- Learning And Electronic Communication
- FARMSCAPE Online Developing A Method For Interactive Internet Support For Farmers Situated Learning And Planning 1
- Word To Web Publishing For Agricultural Research 1
- E Business – What Does It Offer Farm Management 1
- Promotion And Teaching Of Agricultural Science And Languages In Schools A Practical Approach 1
- Co Ownership Of Agricultural Research Data – A Difficult Issue In The Digital Age 1
- Internet Technology For On Farm Self Directed Learning In South Queensland 1
- Farming Systems 4: Water In The Landscape
- Soil Moisture And Canola Yield In An Alley Farming System 1
- Solving The Dryland Salinity Problems Of The Murray Darling Basin With Historical Research, Observation And Logic 1
- Incorporating Lucerne Leys Into Cropping Systems On The Clay Soils Of The Darling Downs 1
- Comparing Newly Established Lucerne With Annual Pastures In South East Western Australia 1
- Real Time Analysis Of Rainfall, Soil Water Content And Surface Runoff 1
- Phase Farming Grains With Lucerne In South East Australia 1
- Break Crops: Legumes And Other
- Utilising The Full Yield Potential Of New Pulse Cultivars In Victoria Through Improved Agronomy 1
- Progress Towards Reducing Seed Toxin Levels In Common Vetch (Vicia Sativa L.) 1
- Agronomic And Economic Potential Of Grain Legumes In Tasmania 1
- New Tropical Industrial Hemp 1
- Population And Sowing Depth Effects On Yield Components Of Grain Legumes 1
- Importance Of Seed Discolouration In Faba Beans (Vicia Fabae) Grown In Southern Australia 1
- Learning And Electronic Communication
- Concurrent Session 6, 1030 1200, Thursday 1 February
- Breeding And Selection
- Pollen Ovule Ratios As A Method Of Estimating Breeding System In Trifolium Pasture Species 1
- Improving Grain Yield By Selection For Greater Early Vigour In Wheat 1
- Improving The Efficiency Of Pasture Breeding Programs Through The Use Of Spatial Analysis 1
- Exploring Lucerne Germplasm Diversity For Southern Cropping SYSTEMS 1
- Developing Grazing Tolerant Lucerne 1
- Breeding For Improved Zinc And Manganese Efficiency In Wheat And Barley. 1
- Prime Quality High Protein Wheat Variety Evaluation In South Australia 1
- Developing And Implementing Molecular Markers In Perennial Ryegrass Breeding 1
- Assessment Of Aluminium Stress Tolerance Of Triticale Breeding Lines In Hydroponics 1
- Crop Management, Populations, Canopy Structure
- High Sowing Rates Reduce Seed Weight In Canary Seed (Phalaris Canariensis L.) 1
- Effects Of Plant Population On Pyrethrins Yield Of Pyrethrum, (Tanacetum Cinerariifolium) In Tasmania 1
- Modification Of Within Canopy Microclimate In Maize For Intercropping In The Lowland Tropics 1
- Compensatory And Competitive Ability Of Two Canola Cultivars 1
- Canopy Architecture And Nitrogen Utilisation For Biomass Production: The Contrast Between Maize And Sunflower 1
- Faba Bean Seeding Rates For Central And Southern NSW 1
- Earliness Management Systems For Cotton Production 1
- Unique Root Mass In Australia’s Rice 1
- Achieving The Genetic Potential Of Peanuts In Irrigated Production Systems 1
- Weed And Pest Management
- A Comparison Of Solarisation And Secondary Cultivations To Control Papaver Dubium L. In Commercial Poppy Crops. 1
- Evaluating Potential Strategies For Effective Lucerne Removal Prior To Cropping 1
- Control Of Saffron Thistle By Grazing Management 1
- Manipulating The Composition And Yield Of An Annual Pasture Prior To Cropping. 1
- Imazethapyr Recropping Recommendations For Canola Are Suitable For Australia’s Neutral Alkaline Soils. 1
- Lime And Crop Rotations For Weed Seedbank Management In New Pastures 1
- Organic Weed Management Survey: Methods Used By Australian Herb And Vegetable Growers 1
- Metsulfuron Methyl Residues And Potential Recropping Damage In Victorian Cropping Soils. 1
- The Effect Of Glyphosate On Whitegrass (Cortaderia Pilosa) 1
- Slugs, Snails And Iron Based Baits: An Increasing Problem And A Low Toxic Specific Action Solution 1
- Crop Modelling And Management
- Adapting APSIM Lucerne To The Western Australian Environment 1
- Soil Acidification And Liming In The Low Rainfall Wheatbelt Of South Western NSW. 1
- Simulating Response Of Canola To Sowing Date In Western Australia 1
- TRIAL YEARS Without Tears: Enhancing Recommendations Of Flowering And Yield In Wheat 1
- Land Clearing In The Central Western Wheatbelt Of NSW Affects Soil Acidity 1
- Modelling Water Stress Response In Sugarcane: Validation And Application Of The APSIM Sugarcane Model. 1
- Greencalc: A Calculator For Estimating Greenhouse Gas Emissions For The Australian Sugar Industry 1
- Validation Of The APSIM Lucerne Model For Phenological Development In A Cool Temperate Climate. 1
- Can The APSIM Model Simulate Wheat Yield And Grain Protein In South Western Queensland? 1
- Breeding And Selection
- Conference Information
- Poster Sessions
- Farming Systems
- Nitrogen Fixation Inputs From Lucerne Dominated Pastures In The Central East Of NSW 1
- Grain Sorghum As A Dryland Cropping Option In The Wimmera Region, Victoria 1
- Raised Bed Cropping Leading The Way In High Rainfall Southern Australia 1
- Using Lucerne To Improve The Reliability Of Cropping On Waterlogged Soils 1
- Agronomy And Policy Design: Tackling The ‘where, What And How’ Questions 1
- Simulation Of Tactical Use Of Phase Farming To Reduce Deep Drainage 1
- Soil Water Availability And Root Distribution In Rainfed And Irrigated Lucerne 1
- Adoption, Extension And Education
- Butterfly Pea – A Legume Success Story In Cropping Lands Of Central Queensland 1
- Developing A State Extension Program TOPCROP In Victoria 1
- Challenging Sowing Rates For Wheat To Achieve Target Plant Densities 1
- Enhancing Student Learning Using Decision Support Tools 1
- Management Techniques And Turnip Variety Effects On Turnip Yield In Western Victoria 1
- Land Use
- Disease, Weed And Pest Management, Allelopathy
- Field Selectivity Makes Emamectin Suitable For IPM 1
- Phytotoxicity Of Wheat Leachates And Ferulic Acid To Germination And Radicle Elongation Of Canola 1
- The Potential Of Summer Crops To Affect Weed Growth 1
- Tolerance Of Cotton To Simulated Helicoverpa Damage 1
- Characterisation Of Water Soluble Phytotoxins From Vulpia Spp. Residues 1
- The Effect Of Rutherglen Bug (Nysius Vinitor) On Yield And Quality Of Hybrid Carrot Seed 1
- Annual Ryegrass Control In Conventional And Herbicide Tolerant Canola 1
- Control Of Vulpia Spp. (silvergrass) In Perennial Pastures Of South Western Victoria 1
- Wheat Allelopathic Potential Against A Herbicide Resistant Biotype Of Annual Ryegrass 1
- Crop Agronomy Plant Density, Sowing Date
- Variation In Growth, Yield And Quality Of Wheat Cultivars On The South Coast 1
- Effect Of Plant Density And Sowing Date On Narrow Leaf Lupin Production In The Victorian Mallee 1
- Plant Density And Planting Date Impacts On Pima Cotton Development 1
- Dry Matter And Nitrogen Remobilization Of Rice Genotypes Under Different Transplanting Dates 1
- Seeding Rate And Date Of Sowing Effects On Two Wheat Varieties In The Victorian Mallee 1
- Crop Nutrition
- Crop Water Use Efficiency
- Pasture Physiology And Management
- Root Development In Perennial Ryegrass Under A Range Of Cutting Treatments 1
- Lucerne Roots Growth As A Function Of Summer Watering 1
- Application Of Woodchip Waste To Sown Pastures 1
- Pasture Characteristics Of Irrigated Dairy Farms In Northern Victoria 1
- Effects Of Renovation On Irrigated Dairy Pastures In Northern Victoria 1
- Productivity And Nitrogen Dynamics Of Pasture Under Warmer, High CO2 Conditions 1
- Using Legume Species Mixtures To Increase And Stabilise Legume Content In Pastures 1
- Breaking Dormancy In Yellow Serradella Seed 1
- Seasonal Alkaloid Concentrations In Perennial Ryegrass Dairy Pasture 1
- Shade Adaptation Of Established Turfgrass Improved By Plant Growth Regulator 1
- Ryegrass Cultivar, Lock Up Duration And Nitrogen Effects On Silage Yield And Quality 1
- Effects Of Grazing Method And Soil Fertility On Pasture And Wool Production 1
- Application Of PastureProbe™ In Determination Of Herbage Mass Of Subterranean Clover 1
- Forage Crop Management Water Use
- Farming Systems Other
- Demonstration Of Organic Crop Production At Rutherglen, North East Victoria 1
- Farmer Practices And Attitudes To Conventional Fallow In The Low Rainfall Mallee 1
- Nitrogen Mineralisation From Shoot And Root Residues Of Crop And Pasture Species 1
- Management Effects On Cumulative Dry Matter Production And Nitrogen Uptake In Wheat 1
- Simulation Of Yield And Environmental Impacts Of Wheat After Rice In Bangladesh And Australia 1
- Soil Management Including Species For Difficult Soils
- Effects Of Long Term Subsurface Drip Irrigation On Soil Structure 1
- A Simple Method For Quantifying Effects Of Management Practices On Cracking Clays Under Dryland Conditions 1
- The Performance Of A Range Of Annual Pasture Legumes On Acid Saline Soils In Tasmania 1
- Reactivity Of Southern Murray Mallee Soil Carbonates 1
- Improving The Adaptation, Profitability And Reliability Of Pulses Growing On Hostile Alkaline Subsoils 1
- The Response Of Lucerne (Medicago Sativa L.) To The Combined Effects Of NaCl And P 1
- Productive Reclamation Of Saline Soils With Distichlis Spicata Var. Yensen 4a 1
- Evaluation Of Cotton Transplanting In Salin Soils 1
- Physiology, Crop Models, Decision Support
- Physiology, Agronomy General
- Effects Of Water Stress On The Development Of The Inflorescence In Two Spring Wheat 1
- Conder Broad Adaptation Of A Peanut Variety Selected For Irrigation 1
- Using Canola Monitoring Groups To Understand Factors Affecting Canola Production In Esperance 1
- Rapid Image Analysis For Counting Engorged Pollen Grains Of Rice 1
- Study Of Some Factors Affecting Carrot Taproot Size Uniformity 1
- The Effect Of Various Pre Treatments On Soybean Seed Performance 1
- Growth Attributes For Higher Wheat Yields In The High Rainfall Zone Of South Western Victoria 1
- The Early Growth Of Linola(/linseed (Linum Usitatissimum L.) In Tasmania 1
- Crop Agronomy Breeding
- Pasture Nutrition
- Pastures Breeding And Selection
- Overcoming The Seed Production Barriers With Native Grasses 1
- Species Of Austrodanthonia On An Acid Soil On The Southern Tablelands Of NSW 1
- Performance Of Experimental Synthetic Cultivars Developed From The Kangaroo Valley Ecotype Of Perennial Ryegrass (Lolium Perenne L.) In Australia 1
- LIGULE – The Distribution Of Promising Native Grasses – A Geographic Perspective 1
- Photosynthetic Variation In Genotypes Of Perennial Ryegrass (Lolium Perenne L.) Selected To Map Drought Tolerance 1
- Ryegrass, Rust And Resistance 1
- Identification Of Novel Germplasm To Develop New Tall Fescue Cultivars For Australia 1
- Cultivar And Species Comparisons Pastures
- Legumes For Agroforestry Systems 1
- Variation For Components Of Seedling Vigour Among Persian Clover Cultivars 1
- Drought Resistance Of Native And Introduced Perennial Grass Species 1
- Alternative Perennial Grass Species For North Facing Slopes In Tasmania 1
- The Register Of Australian Herbage Plant Cultivars: Delivering Information For Agriculture 1
- Evaluation Of Annual Medics (Medicago Spp.) In The Victorian Mallee 1
- Lucerne For Dryland Farming Systems In The Queensland Subtropics 1
- Lotus Uliginosus – A Potential Pasture Species For The Falkland Islands 1
- Extending Pasture Quality Later Into The Season 1
- Prairie Grass: An Alternative To Perennial Ryegrass In Subtropical Pastures 1
- Developing Molecular Markers For Traits Associated With Drought Tolerance In Perennial Ryegrass (Lolium Perenne L.) 1
- Farming Systems
- Plenary Papers
- Opening Session
- Technology On The Farm
- Scientific Support For Decision Making
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Imperatives For Australian Agriculture Andrew Campbell
- Understanding Climate Variability To Improve Agricultural Decision Making
- Towards A Common Advisory Toolkit For Managing Temperate Grazing Systems
- Farming Systems Research
- Participatory Research Processes And Results
- Biotechnology, Food And Agronomy
- The Big Picture
- Concurrent Session 1, 1300 1430, Monday, 29/1/2001
- 2003 11th AAC
- Contributed
- Biotechnology
- Lucerne
- Performance Of APSIM Lucerne In South Australia
- Rate Of Root Growth In Lucerne Varies With Soil Type In Western Australia
- Farmers’ Experiences With The Companion Cropping Of Lucerne In North Central Victoria
- Conservative Water Use By Lucerne
- Where Should Lucerne Be Grown In The Western Australian Cropping Zone?
- Attitudes And Needs Of Grain Growers To Developing Lucerne Based Farming Enterprises
- Effect Of Lucerne Density On Soil Moisture Content During Summer In Southern NSW
- New Crops & Crop Products
- Evaluation Of Agronomic Characters Of Mulberry Varieties In South East Queensland
- The Growth And Development Of Industrial Hemp (Cannabis Sativa) In Central Western New South Wales
- Evaluation Of Improved Lines Of Guayule (Parthenium Argentatum Gray) For Commercial Rubber
- The Search For Alternative Safflower (Carthamus Tinctorius L.) Cultivars Adapted To Southeastern Australia With Improved Marketability
- Crambe: A North Dakotan Case Study
- A New Temperate Forage Legume With Great Potential – Breeding New Cultivars Of Hedysarum
- Biopolymers From Crops: Their Potential To Improve The Environment
- Nitrogen In Rotations
- Soil N Mineralisation Following Fallow, Annual Crops And Perennial Pastures
- Silage Cutting And Grass Removal In The Pasture Phase, Their Effect On Two Consecutive Cropping Years
- Simulating Crop And Soil Processes In Crop Sequences In Southern NSW
- Nitrogen Benefits To Wheat From Pasture Legumes In A Low Rainfall Cropping System
- Organics
- Pastures
- Seasonal Changes In Feed Quality Of Dorycnium Spp.
- The Change In The Proportions Of Annual Legume Species In Response To The Presence Of Lucerne And The Addition Of Gypsum.
- Hedysarum, A New Temperate Forage Legume With Great Potential Field Evaluation
- Are Mixtures Of Annual Pasture Legumes More Productive And Persistent Than Monocultures On The Low Rainfall Alkaline Soils Of Southern Australia?
- Effect Of Different Rates Of Dairy Effluent On Turnip DM Yields And Nutritive Characteristics
- Effects Of Pasture Management And Liming History On Pasture And Wheat Performance In The Central Wheatbelt Of Western Australia
- Trifolium Spumosum L. An Exciting Prospective Legume For Fine Textured Soils In Mediterranean Farming Systems
- Effect Of Timing And Intensity Of Drought On The Seed Yield Of White Clover (Trifolium Repens L.).
- Use Of Paired Soil And Plant Tests To Set Soil Fertility Targets For Subterranean Clover Based Pastures
- Comparing Sulfur Sources For Pastures On Light Sandy Soils
- Perennial Ryegrass Survival Through Summer
- Precision Agriculture
- Potential Cost Savings From Variable Rate Technology In Site Specific Crop Management In The Victorian Mallee
- Recurrence Of Yield And Protein Variation In The Northern Grains Region
- A Multivariate Evaluation Of Sugarcane Farm Performance Over Time Based On Yield And Commercial Cane Sugar
- Conceptual Implications For Water Use Of Cereals Derived From Spatial Variability In Yield Maps
- Detecting Subsoil Constraints On Farms In The Murray Mallee
- Determining Zones For Yield Response To N Fertiliser In A Paddock With Subsoil Limitations, Using Electromagnetic Induction Techniques
- Precision Agriculture Solutions To Underpin Profitable Land Use Change For A Better Environment: A Case Study In Western Australia
- Risk Assessment & Management
- Soil Constraints & Management
- Water Use Efficiency Of Wheat In A Semi Arid Environment
- The Role Of ‘boron Tolerance’ In Improving Adaptation Of Cereals To Highly Alkaline Subsoils.
- Yield And Protein Benefits From Application Of Nitrogen Fertiliser To Wheat On Upper Eyre Peninsula
- Phosphorus – Yield Responses In Wheat, Canola And Field Peas Grown At Different Soil Colwell P And PBI Levels.
- Salt Tolerant Lentils – A Possibility For The Future?
- Lime Response Of Winter Crops On A Duplex Soil At Oolong In SE NSW
- The Effects Of Salinity, Sodicity And Soluble Boron On Wheat Yields In The Victorian Southern Mallee.
- Weeds
- Weed Interference In Soybean (Glycine Max)
- Trends In Yielding Ability And Weed Competitiveness Of Australian Wheat Cultivars
- Rethinking The Management Of Serrated Tussock, Our Worst Perennial Grass Weed
- Development Of A Parasite Module In APSIM Case Study: The Parasitic Weed Orobanche Crenata Infesting Faba Bean
- Working With Communities
- Crop Agronomy Modelling
- The Impact Of Crop Modelling On Plant Physiological Research And Breeding – An Example.
- Variability In Lupin Yield Due To Climate In Western Australia
- On Farm Testing Of The Sirius Wheat Calculator For N Fertiliser And Irrigation Management
- Adaptation Of The APSIM Wheat Module To Simulate The Growth And Production Of Wheat On Hostile Soils
- Crop Agronomy Monitoring
- Crop Agronomy Pulses & Oilseeds
- The Response Of Lentil Cultivars To Sowing Date And Plant Density In The Southern Mallee Of Victoria
- Poor Growth Of Canola In Retained Wheat Straw – Causes And Consequences
- Matching Lupin Cultivar To Environment In Western Australia
- Intercropping With Canola Improves The Productivity And Sustainability Of Field Pea
- Developing Protocols To Quantify Biomass In Commercial Faba Bean Crops
- Yield And Seed Discolouration Of Windrowed Broad Beans In Victoria
- Variation In Yield Of Faba Bean Across Southern Australia Following Rhizobial Inoculation
- The Role Of Phenology In Adaptation Of Chickpea To Drought
- The Comparative Growth, Yield And Water Use Of Safflower, Linola™, Mustard, Canola And Wheat In Southern Australia
- Crop Agronomy Warm Season
- Cold Shock In Early Growth Of Cotton
- The Impact Of Crop Rotation On Peanut Productivity In Rainfed Cropping Systems
- Avoiding Low Temperature Damage In Australia’s Rice Industry With Photoperiod Sensitive Cultivars
- Canopy Development In Phosphorus Deficient Sweet Corn.
- Intensity Of The Production System Influences The Impact Of Yield Decline In Sugarcane
- Assessing The Likelihood Of The Occurrence Of Low Temperature Damage In The NSW Rice Industry
- Evaluation Of Sorghum In Western Australia
- Optimisation Of Nitrogen Supply From Sugarcane Residues In The Wet Tropics
- Determinacy In Cotton: Measurement And Potential Implications
- Crop Growth And Maturity In Ultra Narrow Row And Conventionally Spaced Cotton
- Accounting For Cotton Water Use And Productivity: A Tool To Estimate On Farm Water Use Efficiencies
- Benefits Of Sub Surface Application Of Nitrogen And Water To Trickle Irrigated Sugarcane
- Crop Agronomy Wheat
- Yield And Falling Numbers Of New Wheat Cultivars On The South Coast Of Western Australia
- Breeding Wheat Lines For Southern Australia According To Three Climatic Mega Environments
- Effect Of Previous Crops On Crown Rot Infection And Yield Of Wheat
- Durum Wheat A Profitable Crop For Western Australia
- Successful High Rainfall Cropping In Southern Australia Using Raised Beds
- Poor Wheat Crops Following Canola A Survey Of Farmers And Agronomists
- Necrotic Tipping And Yellowing Of Leaves In Wheat (cv. Diamondbird) In Southern New South Wales In 2001
- The Influence Of Climatic Factors And Crop Nutrition On Seed Vigour In Wheat
- Enhanced Tolerance Of High P Plants To Environmental Stresses Is Related To Primary Root Diameter And Potential Root Hydraulic Conductivity For Water And Nutrient Uptake
- Water Soluble Carbohydrates And Yield In Wheat
- Crop Yield Response To Pasture Legumes In A Pasture Crop Rotation
- Crop Water Loss
- Environmental Management
- Controlling Nitrogen Losses Can We Learn From The European Experience?
- Reducing Deep Drainage Through Controlled Runoff Management In High Recharge Tablelands Landscape
- Botanical Diversity Within Two Saline Ecosystems In Southwestern Australia
- Environmental Management Systems – New Tools For Plant Based Solutions To Salinity
- How Can Farmers And Researchers Measure Sustainability?
- Management Options To Minimise Atrazine And Metolachlor Export From Grain Cropping Areas
- Ecological And Productivity Potential Of Native Woody Species In Australian Cropping Systems
- Farming Systems
- Long Term Farming Systems Analysis In The Southern Mallee And Northern Wimmera.
- The Effect Of Row Configuration On Yield Reliability In Grain Sorghum: II. Modelling The Effects Of Row Configuration
- The Effect Of Row Configuration On Yield Reliability In Grain Sorghum: 1. Yield, Water Use Efficiency And Soil Water Extraction
- Donald Medal Oration
- Invited
- Biotechnology
- Global Warming
- Global Environmental Change And Future Crop Production
- Risk Analysis Of Possible Environmental Change And Future Crop Production In South Australia
- Identifying On Farm Management Practises Aimed At Reducing Greenhouse Gas Emissions From Sugarcane Primary Production
- National And Regional Assessments Of Crop Yield Trends And Water Use Efficiencies
- Marketing
- New Crops
- New Solutions For Salinity
- Linking Farm Management With Catchment Response In A Modelling Framework
- Solutions For The Environment – But What Are The Problems? An Exploration Of Approaches To Deal With Issues Of The Environment
- A Revolution In Agriculture Is Needed If We Are To Manage Dryland Salinity And Related Natural Resource Issues
- Non Fallow And Tillage Options For Control Of Dryland Salinity In The Murray Mallee
- The Impact Of A Lucerne Phase In A Crop Rotation On Groundwater Recharge In South West Australia
- Precision Agriculture
- Conference Information
- Poster
- Biotechnology
- New Solutions For Salinity
- Deep Drainage Under Cropping In Western Australia
- Developing Sustainable Farming Systems With The Aid Of Precision Agriculture Tools
- The Effect Of Saline Irrigation Water On Perennial Pasture Quality In The Goulburn Valley
- Identifying And Developing Forage Resources For Saline Discharge Areas.
- Production Of Forage From Saline Water
- Organics
- Pastures
- Root Biomass Production, Root Distribution, And Soil Water Dynamics Of Three Alternative Perennial Pasture Legumes In Comparison With Medicago Sativa During Their Early Growth
- Thermal Time Requirements For Seedling Development Of Caucasian And White Clovers
- Establishing Grass Legume Pastures On Rundown Cropping Soils Of The Western Downs In Southern Queensland.
- An Evaluation Of Tropical Pasture Legumes On Gidgee Soils In The Semi Arid Tropics
- Growth And Water Use Of Perennial Ryegrass And Lucerne In Summer
- Phosphorus Requirement Of Pasture Species And The Influence On Botanical Composition.
- Plant Species Diversity And Productivity In Grazed Permanent Grasslands
- Impact Of Renovation On Production Of Irrigated Pasture In Northern Victoria
- Factors Affecting Phalaris Persistence
- Long Term Effects Of Nitrogen Fertiliser On Nitrogen Fixation In Grazed Perennial Ryegrass / White Clover Dairy Pastures In South West Victoria
- Medics As Self Regenerating Cover Crops In Canadian Prairie Grain Systems
- Glycinebetaine Foliar Application Increases Pasture Winter Growth And Milk Yield
- Pasture And Soil Effects Of Superphosphate Under Cell Grazing Management
- Estimating Plant Density Using A Modified Carter Ring Method
- Development Of Near Infrared (NIR) Spectroscopy Techniques For Analysing The Nutritive Value Of Fresh Silage.
- Simulated Time Of Break And Emergence Of Annual Species In A Degraded Perennial Grass Pasture
- Impacts Of Sub Surface Drainage And "on Off" Grazing In Reducing Wet Soil Pugging Damage On Southern Victorian Dairy Pastures Pasture Effects
- Impacts Of Sub Surface Drainage And "on Off" Grazing In Reducing Wet Soil Pugging Damage On Southern Victorian Dairy Pastures – Some Soil Effects
- Effects Of Fungal Endophyte On The Persistence And Productivity Of Tall Fescue At 3 Sites In Eastern Australia
- Clover Species Suiting Crop Rotations
- Precision Agriculture
- Risk Assessment & Management
- Soil Constraints & Management
- Crop Agronomy Monitoring
- Crop Agronomy Pulses & Oilseeds
- The Effect Of Row And Plant Spacings On The Growth And Yield Of Chickpea (Cicer Arietinum L.)
- The Effect Of Sowing Date And Rate On Seed Coat Discolouration Due To Frost In Field Peas In The Southern Mallee Of Victoria.
- Effect Of Seeding Rate On Organic Canola Emergence, Red Legged Earthmite (Halotydeus Destructor) Density, Weed Biomass And Grain Yield In Northeast Victoria.
- Assessment Of Possible Allelopathic Interactions Between Soybean (Glycine Max) And Amaranthus Powellii And Cyperus Rotundus Using In Vitro Systems.
- Optimum Plant Densities For Faba Bean Cv Fiesta VF Sown On Raised Beds
- The Effect Of Sowing Date On Yield Of Different Lentil Types
- Turnip Dry Matter Yield And Water Use Efficiency Under Different Irrigation Regimes In Western Victoria
- Limits To Achieving Potential Yield Of Canola In Southern NSW
- Legume Shoot And Root N Accumulation Under Water Stress
- Genotype X Environment Interaction On Seed Yield Of Indian Mustard (Brassica Juncea L.) And Canola (Brassica Napus L.) In A Mediterranean Type Environment Of South Western Australia
- Effects Of Water Stress On Water Relations And Yield Of Indian Mustard (Brassica Juncea L.) And Canola (Brassica Napus L.)
- Control Of Ascochyta Blight In Chickpeas Using Resistant Varieties And Foliar Fungicides
- Development Of A Soil Assay For Screening Rapeseed (Brassica Napus L.) Resistant To High Manganese
- Effect Of Large Seed Size, Post Sowing Compaction And Chemical Seed Dressings On The Survival Of Canola Seedlings In The Presence Of The Earth Mite Damage
- Performance Of Brassica Genotypes In Contrasting Environments
- Crop Agronomy Warm Season
- Dry Season Irrigated Rice Yield Response To Time Of Sowing In Laos
- Root Health And Nutrient Accumulation In Sugarcane.
- Physiological Responses Of Cotton To Subsurface Drip Irrigation On Heavy Clay Soil
- Genetic Variation For Post Anthesis Drought Resistance Traits In Grain Sorghum
- Stem Diameter: A Rapid And Accurate Parameter For Monitoring Growth Of Sorghum
- Rotation And Biocide Effects On The Growth And Yield Of Sugarcane
- APSIM Rice: A Growth And Development Model For Rice
- Effects Of Cutting And Sowing Date On Biomass Production And Nitrogen Content Of Forage Sorghum
- Impact Of Crop Management On Cotton Crop Maturity And Yield
- Modifying Sugarcane Irrigation To Account For A Shallow Water Table
- An Evaluation Of Safflower, Linola™, Sunflower, Maize, Buckwheat And Sorghum As Spring Sown Cropping Options For South Eastern Australia
- The Performance Of Imported Safflower (Carthamus Tinctorius L.) Hybrids In South Eastern Australia
- Crop Agronomy Wheat
- Early Vigor Wheat Genotypes Have Better N Uptake
- Partitioning Of Nitrogen Among Monomeric Protein Fractions During Grain Development In Wheat Is Not Influenced By Nitrogen Nutrition And Post Anthesis Temperature
- Optimum Management Of N Fertiliser For Wheat Growing On Alkaline Soils
- Timing Of Late Applications Of N Fertiliser And Season On Grain Yield And Protein In Wheat
- Time Of Sowing Affects Small Grain Screenings In Wheat In A Dry Season
- Screening Of Tolerant And Susceptible Wheat Cultivars To Waterlogging In The High Rainfall Zone Of The Southwest Victoria
- Environmental Management
- Modelling Temperature Effects On Plant Residue Toxicity
- Runoff From Raised Bed Crops In South West Victoria, 2001
- NutriSmart®: A Fertiliser Capable Of Re Establishing The Sustainability Of Ecosystems And Enhancing The Productivity Of Farmland
- Soil Nitrogen Mineralisation As Affected By Size Of Rainfall Events
- Farming Systems
- Variety Interactions With Wheat Row Spacing And Seeding Rate
- Developing A Minimum Till Cropping Package For The Lower North Central Catchment Of Victoria
- Effect Of Cropping Rotation On Weed Incidence And Botanical Composition Of Ensuing Pastures At Oolong In SE NSW
- Interactive Effects Of Drought And N Stress On Wheat And Canola Growth
- Nitrogen And Moisture Regimes For Genotype Differentiation In Breeding For Consistently Low Grain Protein Concentration In Barley.
- A Comparison Of Four Farming Systems In Central Western NSW
- Fertiliser Inputs For Maximum Yield And Quality Of Gairdner Barley
- Within Field Protein Variation In The Northern Grains Region
- Evaluation Of A New Class Of Fungicides On Grain Yield In Wheat And Barley
- Simulating Dynamics Of Allelochemical Production From Living Plants
- Grain Grower Perceptions Of The Economic Value And Longevity Of Glyphosate
- Have We Found A New Sustainable Farming System?
- Estimating The Optimal Relative Density Combination Of Two Crops In An Intercrop
- Detection Of Elemental Response With The Use Of Fertiliser Test Strips
- Farming Systems Research In Australia; Origins And Integrations
- Mycorrhizae Reduce Carbohydrate Reserves Of Wheat
- Pasture Composition And Timing Of Removal – Effects On Wheat Yield And Protein Level.
- Effect Of Waterlogging On The Growth Of Barley
- Lucerne
- New Crops & Crop Products
- Variation For Osmotic Adjustment In Australian Triticale Cultivars
- Seed Softening In Hedysarum Spp. – New Temperate Forage Legumes With Great Potential
- Flowering And Seed Development Of Torilis Nodosa And Anthriscus Caucalis.
- An Examination Of The Seedbank Distribution, Seedling Emergence And Seed Survival Of Apiaceae Weeds In Pyrethrum
- Flowering Characteristics Of Adzuki Bean From China
- Contributed
- 2004 12th AAC, 4th ICSC
- Copyright And Citation
- Donald Oration
- Conference Information
- About This Publication
- Asian Crop Science Association
- The Australian Society Of Agronomy
- Committees And Reviewers
- Media Releases
- Global Agriculture At The Cross Roads
- Support For Science & Technology Essential To World Agriculture And Trade
- Wheat Monoculture Is Sustainable
- Fine Tuning Improves Water Productivity
- To Save Water, Consume Less Meat?
- The World Water Crisis: Fact Or Fiction?
- Feeding The Future: Plant Genetic Resources As Global Public Goods
- Australia Leads Crop Water Use Research
- Novel Tillage Practice To Increase Yields
- Zero Tillage: A Blessing In Disguise? Herbicide Resistance Looms As Major Threat
- Key Wheat & Maize Scientists Attend Crop Science Congress In Australia
- Radical Surgery Is The Answer To Innovation Woes
- Organic Farming Ideology Limits Sustainability
- The Westernisation Of Asian Diets
- Plants As Bio Factories
- New Options Coming For Salinity Management
- GMO’s – Case By Case, Country By Country
- Australia Supports Research On Crop Diversity
- Conference Programs
- Plenary Speakers
- Sponsors
- Welcome
- Plenary Papers
- Crop Science, Poverty And The Family Farm In A Globalising World
- Science And Technology: The Essential Way To Keep Sustainable Development Of World Agriculture And Trade
- Stocktake On Cropping And Crop Science For A Diverse Planet.
- Theme 0. Crop Science For A Diverse Planet
- 1. Crop Science For Addressing Water Scarcity
- 2. Crop Science For A Sustainable Future
- 3. Crop Science For Harnessing Genetics
- 4. Crop Science For Effecting Change
- Poster Papers
- Forecasting Trend Of Rice Production Of The World And Regions
- WTO And Internal Factors Which Impact On The Oilseed Economy Of India
- A Long Term View Of Australian Agriculture: History, Future And Lessons
- Diversity Of Grain Cropping In Australia As An Indicator Of Sustainability
- 1. Water
- Breeding Crops That Do Better Under Water Stress
- Heritability Of Drought–Resistance Traits In Peanut
- Morpho Agronomical Variation Among Diverse Lentil Genotypes Under Rainfed Environments In Nepal
- Enhancing Adaptation Of Large Seeded Kabuli Chickpea To Drought Prone Environments
- Genotypic Differences For Drought Resistance In Lablab Purpureus L.
- 1354_welckerc
- Identification Of Drought Tolerant Sweet Potato (Ipomoea Batatas (L.) Lam) Cultivars
- Effect Of Drought Tolerance On Preharvest Aflatoxin Contamination In Peanut
- Moisture Stress Studies In Different Chickpea Types
- Two New Food Hull Less Barley Varieties For Rainfed In Egypt
- Amalgamation Of Drought Adaptation And High Productivity In Arid Zone Pearl Millet (Pennisetum Glaucum)
- Screening For Late Season Drought Tolerance In Wheat Genotypes Grown In Iran
- Improving Water Use Efficiency And Drought Tolerance In Groundnut By Trait Based Breeding Programmes In India
- Enhancing Water Productivity In Irrigated Rice
- Adaptation Of Photosynthetic Components Of Chickpea To Water Stress
- Wild Helianthus Anomalus And H. Deserticola From The Desert Southwest USA: A Potential Source Of Stress Genes For Cultivated Sunflower
- Genetics For Improving Crop Water Use Efficiency
- Increasing Crop Water Productivity Through Genetic Improvement For Tolerance To Water Stresses In Maize (Zea Mays L.)
- Recent Advances In The Marker Assisted Selection For Drought Tolerance In Pearl Millet
- A Midwestern USA Perspective On Water Use Efficiency And Drought Tolerance In The Soybean
- Developing High Yielding Wheat For Water Limited Environments In Northern Australia
- Differences In Drought Avoidance Root Characteristics Among Several Millet Species
- Irrigated Rice Based Systems 5ACSC
- Bed Planting: A New Technique To Diversify/intensify Rice Wheat System In India
- 1046_castanedaa
- Comparison Of Productivity Of System Of Rice Intensification And Conventional Rice Farming Systems In The Dry Zone Region Of Sri Lanka
- Agronomic Performance Of Tropical Aerobic, Irrigated, And Traditional Upland Rice Varieties In Three Hydrological Environments At IRRI
- Study On Permanent Bed Planting With Double Zero Tillage For Rice And Wheat In Sichuan Basin
- Planting Pattern Affects Root System Development, Resistance To Water Transport, And Afternoon Depression In Photosynthesis In Rice Plants
- Increasing Water Productivity And Growth Of Rice With Less Irrigation Water
- The Water Balance On Sloping Land In Rainfed Lowland Rice Ecosystem
- Improving Crop And Water Productivity Of Rice Wheat System In Punjab, Pakistan
- Addressing Water Scarcity In Dry Seeded Lowland Rice
- Reducing The Impact Of Muddy Water On Rice Crop Establishment
- Growing Rice On Raised Beds In South Eastern Australia.
- Increasing Rainfed Rice Productivity In Central Java, Indonesia: A Modeling Approach
- Performance Of Rice Cultivars Under Different Resource Conservation Techniques
- Water Productivity Of Rice Crop In Irrigated Areas
- Performance Of Wheat On Beds And Flats In Punjab, India
- Use Of Raised Beds For Increasing Wheat Production In Rice Wheat Cropping Systems.
- Physiology And Improving Productivity Under Water Scarcity
- Root Specific
- Developing Testable Hypotheses On The Impacts Of Sub Soil Constraints On Crops And Croplands Using The Cropping Systems Simulator APSIM.
- Soil Exploration By Sorghum Root Systems In Wide Row Cropping Systems
- Effects Of Chemical Subsoil Constraints On Lower Limit Of Plant Available Water For Crops Grown In Southwest Queensland
- Phosphorus Nutrition Affects Root Morphology Response To Water Deficit At Different Reproductive Stages In An Early Soybean Cultivar
- Effect Of Root Damage Due To Simulated And Real Whitegrub Attack On Physio Biochemical Characters Of Groundnut Under Various Soil Moisture Levels
- 354_durandjl
- Effect Of Root Exudates On Drought And Aflatoxin Resistance Of Peanut Genotypes
- Root Traits Of Different Crops Under Rainfed Conditions In The High Barind Tract, Bangladesh
- Effects Of Mepiquat Chloride On Lateral Roots Initiation Of Cotton Seedling And Its Mechanism
- Penetration Of Hardpans By Roots Of Wheat Under Drought
- Effect Of Semi Dwarf Genes Ton The Root Penetration Ability Ofin Wheat
- Root Growth Of Canola And Wheat In The Deep Yellow Sands Of The Northern Sandplain Of The Western Australian Wheat Belt.
- Development Of The Root System Ofin Peanut (Arachis Hypogaea L.) Analyzed By The Root Box Method
- WUE And Drought Resistance Models
- 1240_farreiok
- Water Saving Strategies In Rain Fed Farming Systems
- Transpiration Efficiency In A Segregating Population Of Sunflower: Inheritance, Correlation With Other Traits And Association With Hybrid Grain Yield
- Improving Harvest Index Is An Effective Way To Increase Crop Water Use Efficiency
- Variation In Water Use Efficiency Of Peanut Varieties Across Peanut Production Regions
- Traits For Improved Drought Resistance Of Winter Wheat In The UK
- Variation In Shoot And Root Growth In Primary Synthetic Wheats Implications For Overcoming Water Deficits In Marginal Environments
- Drought Effect On Water Use Efficiency Of Berseem Clover At Various Growth Stages
- Drought Resistance Of NERICA (New Rice For Africa) Compared With Oryza Sativa L. And Millet Evaluated By Stomatal Conductance And Soil Water Content
- Using A Crop Growth Model To Hypothesize Genetic Traits To Improve Peanut Yield Under Water Limited Environments
- Effects Of Water And Salt Stress On Yield And Components
- Genotypic Variations Of δ 13C In Rice (Oryza. Sativa L. And Oryza. Glaberrima Steud.) In Relation To Transpiration Efficiency And Biomass Production As Affected By Soil Water Conditions And N
- Water And High Temperature Stress Effects On Maize Production
- Study Of Water Stress Effects In Different Growth Stages On Yield And Yield Components Of Different Rice (Oryza Sativa L.) Cultivars
- Response Of Mustard And Canola Genotypes To Soil Moisture Stress During The Post Flowering Period
- Effect Of Water Deficit On Seedling, Plantlets And Compatible Solutes Of Forage Sorghum Cv. Speedfeed
- Determination Of The Most Sensitive Developmental Period Of Wheat (Triticum Aestivum) To Salt Stress To Optimize Saline Water Utilization
- Effect Of Adaptive Metabolites On Water Relations Of Doubled Haploid Wheat Lines At Different Water Levels
- Response Of Chickpea To Short Periods Of High Temperature And Water Stress At Different Developmental Stages
- Changes In Photosynthetic Gas Exchange, Chlorophyll Fluorescence, And Stem Diameter Of Soybean Plants Under Drought Stress Condition.
- Effects Of Water Supply And Light Intensity On The Growth Of Spring Wheat
- Other Physiology And Water Stress
- Water Consumption In Different Heat Tolerant Cultivars Of Snap Bean (Phaseolus Vulgaris L.)
- Compensative Effects Of Chemical Regulation On Physiological Damage Caused By Water Deficiency During Filling Stage Of Wheat
- Will Modifying Plant Ethylene Status Improve Plant Productivity In Water Limited Environments ?
- Use Of Reflectance Measurements To Clearly Identify Water Stress In Wheat (Triticum Aestivum L.)
- Effects Of Plant Growth Regulators On Water Deficit Induced Yield Loss In Soybean
- Antioxidative Enzymes In Sunflower Subjected To Drought Stress
- Leaflet Orientation And Fibrous Root Trait Combination Effects On Water Use Characteristics In Soybeans (Glycine Max) Via Reciprocal Grafts
- Development And Characterization Of RILs For Molecular Mapping Of Reproductive Stage Moisture Stress Tolerance In Rice
- Root Specific
- Manipulating Dryland Systems
- Fluctuations In Spatial Variability Of Wheat Yield
- Optimum Sowing Time For Rainfed Safflower In Southern Australia Is Affected By Soil Water Availability
- Soil Water Availability Determines Optimum Sowing Rates For Safflower In Southern Australia
- Agronomic Responses Of Six Barley Varieties To Different Plant Populations An Old Question Revisited
- Skip Sorghum When And Where Should It Be Used?
- The Effect Seed Source, Size And Frost Damage On Emergence And Grain Yield In Field Pea (Pisum Sativum) Cv. Kaspa
- Capture And Use Of Water By Wheat And Chickpea In Sole Crops And Intercrops, Under Dryland Conditions Of South Australia
- Raised Bed Cropping In Southern Victoria – A Snapshot Of A Productive And Sustainable Option For Waterlogging Prone Soils
- Enhancing Grain Yield Of Rice (Oryza Sativa L.) Under Upland Conditions In Japan
- Comparison Of Measured Changes In Seasonal Soil Water Content Under Rainfed Maize Bean Intercrop Within A Semi Arid Area
- On Farm Monitoring And Management Of Aflatoxin Contamination In Australian Peanuts
- New Imperatives For Irrigated Agriculture
- IMPASSE: A DSS For Assessing Impact Of Saline And Non Saline Irrigation Waters On Crop Yield Reductions
- Study Of Water Use Efficiency, Quantityield And Quality Of Two Sugar Beet Varieties In Line Source Sprinkler Irrigation
- Effects Of Water Supply And Sowing Date On Water Use Efficiency Of Anise (Pimpinella Anisum L.)
- Soybean Crop Production Under Environmental Stress Conditions At Toshky / Egypt
- Crop Production And Drainage In Conjunction With Subsoiling Role In Deteriorated Egyptian Soil
- Controlled Deficit Irrigation Of Alfalfa (Medicago Sativa): A Strategy For Addressing Water Scarcity In California
- The Effects Of Deficit Irrigation On Water Use Efficiency, Yield, And Quality Of Forage Pearl Millet
- Yield And Yield Components Of Dryland Wheat Genotypes Under Supplemental Irrigation.
- A Comparative Analysis Of Crop Water Productivity Of Rice Wheat And Cotton Wheat Rotations In Rechna Doab, Punjab, Pakistan
- Effect Of Dairy Effluent On DM Yields And Nutritive Characteristics Of Perennial Pasture In Late Spring And Summer
- Evaluation Of The Water Use Efficiency Of Dairy Production Using Crops And Pastures
- Water Use Efficiency Of Irrigated Forage Systems In Northern Victoria
- Improvement Of Water Productivity In Surface Irrigation System By Changing The Plant Spacing In Sugar Beet Cultivation
- Water Use Efficiency Response Of Field Grown Muskmelon And Pepper To Environmental Water Status
- Effect Of Irrigation Strategies On Dry Matter Yields And Water Use Efficiency Of A Range Of Forage Species
- Effect Of Different Rates Of Dairy Effluent On Millet Dry Matter Yields And Nutritive Characteristics
- Water Use Efficiency In Sugar Beet, Subjected To Different Sowing Times And Irrigation Regimes In A Mediterranean Environment
- A Simple Versatile Model Of Crop Yield Response To Water Deficit
- Response Of Canopy Architecture, Yield And Water Use Efficiency Of Winter Wheat To Irrigation Time
- Application Of Remote Sensing Technologies To Improve Yield And Water Use Efficiency In Irrigated Peanuts
- Identifying The Potential To Apply Deficit Irrigation Strategies In Cotton Using Large Mobile Irrigation Machines
- Soil Moisture Based Nutrient Management Of Red Pepper
- Effects Of Irrigation Scheduling On Soybean Growth And Yield.
- Growth Responses To Alternative Irrigation Water For Drought Season In Paddy Rice
- Influences Of Chinese Cabbage Growth And Soil Salinity To Alternative Irrigation Waters
- Management For Ultra Early Cotton Systems
- Environmental Consequences Of Water Management In Farming Systems
- Managing The Interface Of Trees And Crops – Trade Offs In Low Rainfall Zones
- Impact Of Alternative Land Use Patterns On Plant Water Use, Surface Water Flow And Drainage On A Topographic Sequence
- Study On Soil And Nutrients Loss With Soil Textures And Two Crops During Rainfall
- Simulation Of Deep Drainage Under A 13 Year Crop Sequence In Southern NSW
- Lost And Reclaimed: A Case Study Of Gully Rehabilitation In Central Kenya Highlands Using Low Cost Measures
- Environmental Impact Assessment Of Surface Run Off To Small Stream In A Steep Cornfield
- Breeding Crops That Do Better Under Water Stress
- 2. Sustainability
- Farming And Cropping Systems
- General Farming System/rotation
- Productivity Of Wheat And Canola In Rotation With Lucerne/sub Clover Or Sub Clover Pasture
- A Strategic Approach To Incorporating Non Cereal Crops In Low Rainfall Regions Of Southern Australia: A Preliminary Analysis Of Probable Yields
- Simulation Of Crop Growth And Soil Water For Different Cropping Systems In The Gansu Loess Plateau, China Using APSIM
- An Integrated Modelling Approach To Enhance Bali Cattle Production In The Mixed Crop/livestock Systems Of Indonesia
- Broadleaf Crops For Diversification Of Dryland Cropping In The Northern Wheat Belt Of Eastern Australia
- New Agro Techniques In Intensive Crop Rotations Under Marginal Conditions Of Upper Egypt
- Systems Modeling And Farmers’ Participatory Evaluation Of Cropping Options To Diversify Peanut Systems In Anantapur Region, India. I: APSIM Simulations To Analyze Constraints And Opportunities
- Systems Modeling And Farmers’ Participatory Evaluation Of Cropping Options To Diversify Peanut Systems In Anantapur Region, India II. Farmers’ Participatory Field Assessment Of Simulated Peanut Systems
- Diversification Of Farming System By Combining Commercial Tree Species With Medicinal Plants To Boost Economy Of Farmers Under Rainfed Conditions
- Pattern And Factor Analyses Of Diverse Plant And Yield Attributes’ Responses To Alternative Crop Rotations And Management Practices
- Improved Short Fallows: Impact On Weed Populations And Maize Growth In Humid Tropics Of Asia
- The Role Of The Universities Of Wyoming And Nebraska In The Development Of Alternative Crops For The US High Plains.
- 595_greefm
- The Whole Farm Impact Of Including Dual Purpose Winter Wheat And Forage Brassica Crops In A Grazing System: A Simulation Analysis
- Integration Of A Brassica Forage Crop Into A Winter Wheat/pasture Rotation
- SIMBA: A Comprehensive Model For Agro Ecological Assessment And Prototyping Of Banana Based Cropping Systems. An Application In The French West Indies
- Improving Farm Productivity Through Crop Intensification In A Crop Animal System Under Rainfed Environment
- Simulating Spatial Issues In Farming Systems With APSIM
- 960_cavigliaop
- 961_rogetsd
- Legumes In Farming Systems
- Grain Yield And Symbiotic Activity Of Cowpea Cultivars Grown In Sole And Intercropping Systems With Maize In The Limpopo Province Of South Africa
- Exploring Cropping Options With Crop Models: A Case Study Of Pigeonpea Versus Peanut In Rainfed Tracts Of Semiarid Southern India
- Legume Maize Association Influences Crop Characteristics And Yields
- Managing Economic And Environmental Sustainability With Better P Nutrition And Summer Forage Legume In Rotation With Wheat In Southern Queensland
- Legumes For Cropping Systems In The Democratic Peoples Republic Of Korea
- Selecting Suitable Pasture Legume Species As Fodder Crops For The Central Wheatbelt Of Southwest Western Australia
- Evaluation Of Purple Vetch (Vicia Benghalensis) As A Green Manure Legume For Irish Potato Production In Matanya, Central Rift, Kenya.
- Common Vetch In Rotation With Barley: A Sustainable Farming System For A Cool, Semi Arid Mediterranean Area
- Response Of Durum Wheat Genotypes To Previous Crop And N Fertilization Under Mediterranean Conditions
- Genetic Improvement Of Strand Medic (Medicago Littoralis Rohde Ex Lois.) For Australian Farming Systems
- Productivity Improvement In Upland Rice Through The Inclusion Of Legumes, In The Far Western Mid Hill Areas Of Nepal
- 955_assengs
- Other
- Development Of Canaryseed Production In The Central High Plains Of The United States.
- Impact Of Regrowth And Remnant Brigalow (Acacia Harpophylla) Trees On Wheat Production In Southern Queensland, Australia
- Rotation Effects And Profitability Of Safflower Compared To Wheat And Oilseeds In Southern Australia
- 1311_diedhioui
- Long Term Effect Of Stubble Burning On Soil Carbon Levels And Wheat Yield In Southern New South Wales
- Possibility To Grow Early Maturity Corn Hybrids For Energetically Dense Silage In Latvian Conditions
- Mycorrhizal Colonisation Of Dry Season Cotton Grown On Virgin Soil In Northern Australia
- Crop Production Potential And Constraints In The High Rainfall Zone Of Southwestern Australia: Yield And Yield Components
- Strip Cropping Vegetables
- Soil Physical Properties In The Long Term Field Experiment “eternal Rye” After 120 Years Of Different Fertilization
- General Farming System/rotation
- Conservation And Reduced Tillage Systems
- Potato Mechanization In Iran
- Soil Carbon Sequestration And Mulch Based Cropping In The Cerrado Region Of Brazil
- Better Residue Management For More Sustainable Cropping
- Crop Variety Response In Direct Seed (no Till) And Conventional Tillage
- Trashlines And Runoff, Erosion, And Crop Yields In Semi Arid Eastern Kenya
- Conservation Tillage, Poultry Litter, Cropping System And Cotton Production
- 978_christeno
- Asian Cropping Systems (5ACSC)
- Wheat Seed Quality – A Study On Farmers’ Seed
- Change In Growth And Productivity From Pruning And Skiffing As A Method To Prolong Pruning Cycle
- Soil Management Systems Improve Water Use Efficiency Of Rainfed Rice In The Semi Arid Tropics Of Southern Lombok, Eastern Indonesia
- Rice Root Distribution Under Various Systems Of Soil Management On Rainfed Vertisols In Southern Lombok, Eastern Indonesia
- Productivity Of Multi Crops Sown On Permanent Raised Beds In The Tropics
- Improving Livelihood Of Potato Farmers In Afghanistan
- Crop And Cropping Systems Research In The Central Terai, Nepal
- Intercropping Of Agri/horti Crops With Special Reference To Mandarin (Citrus Reticulata Blanco) In Sikkim (INDIA)
- The System Of Rice Intensification (SRI) For Super High Yields Of Rice In Sichuan Basin
- 322_dibyl
- Palm Pulverisation In Sustainable Oil Palm Replanting
- Combating Hunger In North Korea Through Super Corn Development And Science Based Sustainable Farming System
- Nutrient Reserves And Recycling From Oil Palm Trunks At Replanting
- Impact Of Mulches On Growth And Yields Of Cassava And Sweet Potato In Tropical Asia
- Grain Yield Of Direct Seeded And Transplanted Rice In Rainfed Lowlands Of South East Asia
- Vegetative Growth, Resource Optimisation And N Productivity Of Oil Palm (Elaeis Guineensis Jacq.) As Influenced By Soil And Fertilization
- Effects Of Genotype And Sowing Time On Growth Of Soybean In The Mountain Region Of Northern Vietnam
- A Methodology To Explore Options For Rice Based Farming Systems In A Humid Subtropical Region: A Case Study For Jiangxi, China
- Performance Of Rice On Beds And Puddled Transplanted Flats In Punjab, India
- Mechanized Crop Care In Non Flooded Rice Production In Malaysia
- Evaluation Of, And Yield Gap Analysis In Rice Using, CERES Rice Ver. 4.0 In Northwest India
- Enhancing Food Security And Income Through Integrating An Upland Crop In The Rainfed Cropping Systems In The Coastal Areas In Vietnam
- Megasporocarps Of Azolla And Their Germination In Varied Paddy Soils
- Impact Of Early Transplanting On Tillering And Grain Yield In Irrigated Rice
- Effect Of Seeding Rate On The Growth And Quality Of Rice Seedlings In The Long Mat Seedling Culture System
- Biocide Dependence And Resistance
- Weeds
- Assessing The Short And Long Term Effects Of Parasitic Weed Infestation On The Productivity Of Faba Bean In The Mediterranean Region: Simulation Studies Using APSIM
- A Proteomic Approach To Analysing Rice Allelopathy On Barnyard Grass (Echinochloa Crus Galli L.)
- Tillage And Herbicide Application Affect Weed Control And Performance Of Tomato
- The Effect Of Weeds On Wheat Grain Yield In Limed And Unlimed Soils
- Tillage System Effects On Seedling Recruitment Pattern And Persistence Of Lolium Rigidum Gaudin (annual Ryegrass) Seed Bank
- Weeds, Herbicide Use And Resistance In Rice Fields Of Sri Lanka
- Using Row Spacing To Increase Crop Competition With Weeds
- Weed Sshifts In Gglyphosate Ttolerant Ccrops – Aa Rregional Pperspective
- Economic Assessment Of Manual And Chemical Control Of Thorny Mimosa In Cassava In Nigeria
- Rooting Patterns In Wheat Differing In Vigour Are Related To The Early Uptake Of Nitrogen In Deep Sandy Soils.
- Wild Oat Control And Malt Barley Response To Fenoxaprop And Tralkoxydim Herbicides
- Comparative Growth And Competition Of Wild Radish (Raphanus Raphanistrum L.) And Wheat
- Controlling Invasive Wild Rice With ACCase Inhibiting Herbicides
- Growth And Yyield Response Of ‘Sunrise Solo’ Ppapaya To Wweed Mmanagement Sstrategies
- Yield Penalty Due To Delayed Weed Control In Corn And Soybean
- Genetic Improvements In Yielding Potential And Inter And Intra Specific Competitive Ability Of Iranian Winter Wheat (Triticum Aestivum L.) Cultivars Released During The Past 50 Years
- Diseases
- Prevalence Of Fusarium Crown Rot Pathogens Of Wheat In Southern Queensland And Northern New South Wales
- 1308_aubertotjn
- Growing Healthy Rice Seedlings Through Soil Solarization: A Low Cost Technology For Increasing Rice Productivity And Profitability
- Effect Of Plant Activators On Disease Resistance And Yield In Tomato And Canola
- Control Of Foliar Diseases Of Field Pea In Southern Brazil
- Interaction Between Ascochyta Intensity And Sowing Arrangement With Chickpea
- 829_dilcib
- Insects
- Weeds
- Nutrient Recycling And Balance In Cropping Systems
- Soil N Cycling And Uptake
- Cereal Response To N Fertiliser In Relation To Subsoil Limitations
- Waterlogging And Nitrogen Management For Wheat In High Rainfall Cropping Areas Of Southern Western Australia.
- Effects Of N Fertilizer Rate And Placement On Wheat (Triticum Aestivum) Establishment, Growth And Yield.
- Canola Response To Side Banded N Rate, Source And NBPT Addition
- 2098_alvaa
- Genotypic Effects On Yield, N Uptake, NUTE And NHI Of Spring Wheat
- Efficient N Management Technologies For Irrigated Rice In Asia
- Effect Of N Deficiency On Photoassimilate Partitioning And Rhythmic Changes In Fruit And Stem Diameter Of Tomato (Lycopersicon Esculentum) During Fruit Growth.
- The Yield And Quality Of Wheat Affected By Individual And Combined Application Of Animal Manure And Chemical Fertilizers
- Yield And N Content Response Of Winter Wheat To 2 Fallow Lengths After 4 Years Of Lucerne In The Gansu Loess Plateau, China
- Need Based Fertilizer Nitrogen Management Using Leaf Color Chart In Irrigated Rice In Punjab, India
- Nitrogen Mineralization Potential Of Rice Wheat Soils Amended With Organic Manures And Crop Residues
- Uptake And Partition Of 15N Labeled Fertilizer By Nitrogen Rate And Cultivar In Rice
- Nitrogen Storage And Remobilization In Brassica Napus L. During The Growth Cycle
- Residual Effects Of Different N Fertilizer Treatments On Wheat Growth And Yield
- Yield And Quality Of Two U.S. Red Potatoes: Influence Of Nitrogen Rate And Plant Population.
- Spectral Reflectance Of Rice Leaves And Leaf Color Charts For N Management
- Nitrogen Accumulation And Recovery From Legumes And N Fertilizer In Rice Based Cropping Systems
- 970_bramm
- Variation Of Nitrogen Use Efficiency And Its Relationships With Growth Characteristics In Korean Rice Cultivars
- Soil OM And Use Of Wastes
- Environmental Hazard Of Excess Dunder On Agricultural Land
- Assessment Of Waste For Use On Agricultural Land
- Using Tropical Grasses To Enhance The Efficiency Of Nitrogen Capture From Dairy Effluent
- Can Manure Lift Crop Production In Communal Lands Of Semi Arid Zimbabwe?
- A Study On Dynamics And Models Of N, P, K Absorption For High Yield Cotton
- Plant Species Effects On Soil Carbon Stocks A Pot Study
- Green Manures Stimulate Root Development Of Maize And Mungbean Seedlings
- Conversion Of Kampong Ewa Rice Fields In Langkawi, Malaysia, Into Organic Rice Farming.
- Effects Of Land Application Of Two Different Biosolids On Plant Growth And Nitrogen Mineralisation In A Red Ferrosol Soil In Subtropical Australia
- Decomposition Rates And Nitrogen Release Of Turf Grass Clippings
- Rice Cultivation Using Organic Farming System With Organic Input Materials In Korea
- Author: Please Include A 250 Word Abstract As Per Author Guidelines. In Method, Some More Physical Or Chemical Information On The Compost Would Be Useful; Otherwise How Could Anyone Repeat The Work. Also Hardness (or Compaction) Measurements Mentioned In M
- Nutrient Use Efficiency, General Fertilizer Studies
- Comparison Of Soil Tests With Plant Response To Nutrients In Selected Soils Of Vanuatu
- Reducing Fertilizer Application And Rice Growth And Yield Mapping By Variable Rate Treatment In Paddy Fields
- 1781_christensenlk
- 184_tamuray
- Has Nitrogen Use Efficiency Changed For Barley In Finnish Growing Conditions?
- Phosphorus
- An Evaluation Of The Phosphorus Benefits From Grain Legumes In Rotational Cropping Using 33P Isotopic Dilution
- Phosphorus Management And Availability In Highly Calcareous Soil
- Modelling The Effect Of Phosphorus On Maize Production And Nitrogen Use Efficiency On Smallholder Farms In Sub Humid Zimbabwe
- Grain Legume Crops Increase Soil Phosphorus Availability To Subsequently Grown Wheat
- Phosphorus Turnover Between Rice Crops In The Rainfed Lowlands From Residual P Fertiliser, Rice Straw And Volunteer Pastures
- Phosphorus Application To Maize Cowpea Sequences In Smallholder Farming Systems Of Zimbabwe
- Phosphorus Uptake Of Sesbania Cannabina And Crotalaria Juncea Associated With Root Nodule Bacteria
- Screening For Genotypic Variation In Phosphorus Uptake Efficiency In Cereals On Australian Soils
- Phenotypic Plasticity Of Rice Seedlings: Case Of P Deficiency
- Phosphate Requirements Of Creeping Bent (Agrostis Stolonifera) Putting Green Turf
- Phosphorus Concentration In Soil And In Winter Wheat Plants
- Other Elements: S, Zn, Microelements
- Response Of Wheat To Enriched Seed And Boron
- Growth Promotive Effects Of 5 Aminolevulinic Acid In The Presence Of Micro Elements On Yield In Komatsuna, Brassica Campestris Var.perviridis Under Alkaline Soil Conditions
- Effect Of Potassium Fertilizer On The Yield Of Plantain Melon Intercropped On An Oxic Paleustalf In South Western Nigeria
- On Farm Seed Priming Reduces Risk And Increases Yield In Tropical Crops
- Zinc Efficiency As Related To Distribution And Internal Requirement
- A Survey Of Tropical Species For Boron Retranslocation
- Response To Boron Toxicity In Boron Efficient And Inefficient Wheat Genotypes
- 907_rosolemca
- Inoculants, Rhizobia, Biofertilizers
- BioBoost: A New Sulfur Oxidizing Bacterial Inoculant For Canola
- Response To And Benefits Of Rhizobial Inoculation Of Soybean In The South Of Vietnam
- Optimizing Legume Production Using Sulfur Oxidizing Bacteria And Rhizobium Consortia
- Enhancing Yield Of Onion (Allium Cepa L.) Through Mycorrhizal Inoculant In Meloidogyne Graminicola Infested Soil
- Optimization Of Sesame (Sesamum Indicum L.) Production Through Bio/natural Inputs
- A New Approach For Quantifying The Role Of Legume Nitrogen In Crop Rotation.
- Effects Of Rhizobium Strains On Growth And Nitrogen Fixation Of Annual Medics
- Biomass Production And Nitrogen Accumulation Of Some Dual Purpose Legumes In Limpopo Province, South Africa
- Growth Promoting Effects Of The Diazotroph Azorhizobium Caulinodans On Canadian Wheat Cultivars
- Soil N Cycling And Uptake
- Managing Climate Risk In Cropping Systems
- Integrating Software Tools And Seasonal Climate Outlooks To Optimise Wheat Gross Margin Profits At Merredin, Western Australia.
- Does ENSO Influence The Break Of Season In South Eastern Australia?
- Using The REML Statistical Procedure To Examine Trends Due To Climate Change
- A Statistical Distribution For Modelling Rainfall With Promising Applications In Crop Science
- Likely Impact Of Climate Change On Wheat And Sorghum Production In Central Queensland
- Effects Of A Changing Climate On Wheat Cropping Systems In Northern New South Wales
- Assessment Of Probabilistic Forecast ‘skill’ Using P Values
- Forecasting With The Madden Julian Oscillation And The Applications For Risk Management
- The Use Of Climatic Data Generator To Cope With Daily Climatic Data Scarcity In Simulation Studies
- Applying Climate Information To Enhance Wheat Based Farming In Rain Fed Areas Of Pakistan
- Can Smallholder Farmers Benefit From Seasonal Climate Forecasts?
- The Use Of Global Climate Forcing For Rainfall And Yield Prediction In Indonesia: Case Study At Bandung District
- Seasonal Climate Forecasting Has Economic Value For Farmers In South Eastern Australia
- A Statistical Model For Seasonal Rainfall Forecasting Over The Highlands Of Eritrea
- Agricultural Impacts Of Regional Variability Of The West African Monsoon
- The Characteristics Of Daily And Monthly Rainfall For A Decision Support System
- Spatial Modelling Of Extreme Temperatures In The Southern Mallee Of Victoria
- Climate Information Contributes To Better Water Management Of Irrigated Cropping Systems In Southern India
- The Development Of Combined Weather And Crop Yield Forecasting Systems For The Tropics
- Quantifying Risk For Maize–bean Intercrop Production In The Semi Arid Region Of Southern Africa
- 941_diazambronacgh
- 943_minguezmi
- Raising Yield Potential In Cropping Systems
- Rice
- A New Diagnostic Tool Of Rice Grain Filling And Its Response To Stresses Using Grain Population Weight And Size Distribution
- Variation In Leaf Photosynthesis Among Wild Species In Genus Oryza And Among The Progeny Of Selected Crosses Of The Wild Species With A Rice Cultivar.
- Does Elevated CO2 Concentration Affect Lamina Length Of Rice Cultivars?
- 1186_audebert
- Rice Yield Enhancement By Elevated CO2 Negatively Correlates With Hastened Occurrence Of Heading –Results Form Five Years Chamber Studies
- Study On The Physiological And Morphological Indices Among The Modern And Old Rice (Oriza Sativa L.) Genotypes
- In Situ Staining The Activities Of Starch Synthesis Related Enzymes Of The Rice Grain
- The Effects Of Nitrogen Application And Assimilate Availability On Engorged Pollen Production And Spikelet Sterility In Rice
- Relationship Between Dry Weight And Spikelet Number Of Each Tiller At Heading In Rice Plants
- Low Grain Ripening In The New Plant Type Rice Due To Shortage Of Assimilate Supply
- Sucrose Metabolism Enzymes In Developing Rice Endosperm: Their Relations To Grain Filling Of Rice Cultivars With Extra Heavy Panicles
- Effects Of Combined Application Of Ethephon And Gibberellin On Growth And Nutrient Uptake Of Rice Seedlings Growing Under Direct Seeding Conditions
- Photosynthesis, Growth, And Yield Of BC2F4 Lines Derived From Oryza Sativa And Wild Rice Species, O. Rufipogon
- Cytokinin Affects The Leaf Levels Of Ribulose 1,5 Bisphosphate Carboxylase/oxygenase And The Accumulation And Partitioning Of Nitrogen In Paddy Rice At The Ripening Stage
- Interactive Effects Of Atmospheric Ozone And Carbon Dioxide On Photosynthesis, Dry Matter Production And Yield Of Lowland Rice
- Effects Of Rising Temperature On Growth, Yield And Dry Matter Production Of Rice Grown In The Paddy Field
- Australia: New Screening Method For Cold Tolerance During The Reproductive Stage In Rice
- Spatial Variation Of Carbohydrate Metabolism In Rice Leaf Sheath
- Planting Pattern Affects The Rate Of Photosynthesis During Ripening In Rice Plants
- Yield Advantage Of Hybrid Rice Induced By Its Higher Control In Tiller Emergence
- Response Of Rice Cultivars To Aerobic Condition
- Biomass Partitioning Of Rice Under Free Air CO2 Enrichment (FACE): Lessons For Crop Models
- Maize And Sorghum
- Investigating The Use Of Sweet Sorghum As A Model For Sugar Accumulation In Sugarcane
- Fertiliser P Effects On Biomass Partitioning And Quality Of Sweet Corn In A Cool Temperate Environment.
- Studies On The Effect Of Plant Density On Maize Growth Using The Richards Function
- 602_hirelb
- Is Increased Radiation Use Efficiency In Sorghum Related To Increased Height?
- Effect Of Stem Infused Chemicals On Kernel Yield, Levels Of Enzymes And Chlorophyll Content In Maize (ZEA MAYS L) Varieties
- 872_gambnbl
- Wheat
- Use Of Resources In 4 Cultivars Of Wheat In The High Rainfall Zone Of South Eastern Australia Protein And Water Soluble Carbohydrates.
- Evidence For Excess Photosynthetic Capacity And Sink Limitation To Yield And Biomass In Elite Spring Wheat
- Wheat Sterility Identification Of Probable Causes And Solutions
- Some Aspects Of The Supernumerary Spikelets Formation In Wheat From The Viewpoints Of Physiology And Genetics.
- Canopy Profile Distribution Of Leaf Area, Light And Nitrogen In Some Iranian Winter Wheat Cultivars Released During The Last 50 Years
- Breaking The 15 T/ha Wheat Yield Barrier
- Effect Of Elevated CO2 Concentration On Net Photosynthetic Rate And Dry Matter Production Of Spring Wheat In The Tibet Plateau
- Broad Leaf Crops
- Double Row And Conventional Cotton In Tulare County, California
- Source And Sink Characteristics Of Various Genotypes With Different Boll Weight In Cotton
- Physiological Changes Of Leaves On Early Reproductive Branch Caused By Changing The Ratio Of Source To Sink In Cotton
- Competition Studies In Four Grain Legumes
- Effects Of Carbon Dioxide Enrichment During Different Growth Periods On Flowering, Pod Set And Seed Yield In Soybean
- Simultaneous Ggrowth Of Ppods And Sseeds Sset On Different Racemes In Ssoybean
- Light Quality And CO2 Induced Changes In Saccharide Content Of Strawberry (Fragaria X Ananassa) ‘Red Joy’ Plants In Vitro
- Effects Of Waterlogging On Root System Of Soybean
- Effect Of Duration Of Reproductive Stages On Yield In Soybean
- Ethylene Biosynthesis, Ripening And Senescence Behavior Of Tobacco (Nicotiana Tabacum L.) Leaves
- Response Of Sunflower To Plant Growth Regulators
- Rice
- Simulation Modelling In Crops
- A Potential Yield Model For Forage Brassicas
- 1161_herrmann
- A Cohort Model For Simulating Forage Brassica Crops With Variable Plant Size
- Simulating Growth And Development Of Lowland Rice In APSIM
- Simulating Damage Effects Of Parasitic Weeds In APSIM: A Generic Cohort Based Approach
- A Linked Process Based Model To Study The Interaction Between Puccinia Striiformis And Wheat
- The Machine Recognition Of Wheat Crop Features From Images Based On Back Propagation Neural Network
- REMS – A Research Experiment Management System To Support The Maintenance And Development Of Crop Growth And Development Models.
- SIMBA: A Comprehensive Model For Agro Ecological Assessment And Prototyping Of Banana Based Cropping Systems. An Application In The French West Indies.
- Modelling Soil Water, Carbon And Nitrogen Dynamics In Wheat/chickpea Rotations: Parameterisation And Evaluation Of APSIM In A Mediterranean Environment
- Field Validation Of Empirical Functions Used To Estimate Crop Water Use
- Main Stem Node Appearance Of Lucerne Regrowth In A Temperate Climate
- Proposal For Early Crop Monitoring Using A Photosynthetic Production Index And Remotely Sensed Data
- 346_stöckle
- Modelling Broccoli Development, Yield And Quality
- 475_jamieson
- Using A Handheld Multispectral Radiometer To Forecast Grain Protein In Northern Australia
- Simulating Crop Phenology
- Crop Growth Models Can Effect Change – A Case History
- Simulating Maize Development Using A Nonlinear Temperature Response Model
- CropSyst VB – Simpotato, A Crop Simulation Model For Potato – Based Cropping Systems: II. Evaluation Of Nitrogen Dynamics.
- CropSyst VB – Simpotato, A Crop Simulation Model For Potato Based Cropping Systems: I. Model Development
- Combining Near Infrared Spectroscopy And Infrared Aerial Imagery For Assessment Of Peanut Crop Maturity And Aflatoxin Risk
- Farming And Cropping Systems
- 3. Genetics
- Genome Sequencing
- A Hexaploid Wheat BAC Sequencing And Analysis Of Microcolinearity Between Wheat And Rice At The Rht D1 Locus
- New SSR Markers For Pearl Millet From Data Mining Of Expressed Sequence Tags
- Analysis And Functional Annotation Of Expressed Sequence Tags For Tef [Eragrostis Tef (Zucc) Trotter]
- Cross Taxa Transferability Of Sequence Tagged Microsatellite Site (STMS) Primers From Pulses To Peanut (Arachis Hypogaea L.)
- Construction And Characterization Of A Bacterial Artificial Chromsome Library Of Hexaploid Wheat (Triticum Aestivum L) Genotype Chinese Spring
- Can Genomics Revolutionise Genetics And Breeding In Sugarcane?
- Large Scale Analysis Of Expressed Sequence Tags From Indica Rice
- Peanut Expressed Sequence Tag (EST) Project And The Marker Development For Cultivated Peanut (Arachis Hypogaea)
- Gene To Phenotype Complexity
- Gene Discovery
- High Nitrogen Conditions Enhance The Cooling Damage To Pollen In Rice Plants : Proteome Analysis Of Mature Anthers
- Cloning And Characterization Of Mitochondrial ATP Synthase 6kDa Subunit Gene In Rice (Oryza Sativa. L)
- Why Is Transferring Apomixis To Crops Still A Dream?
- Identification And Characterisation Of Differentially Expressed Genes In Wheat Undergoing Gradual Water Deficit Stress.
- Identification Of Genes Contributing To High Sucrose Accumulation In Sugarcane
- Exploring A New Detergent Inducible Promoter Active In Higher Plants And Its Potential Biotechnological Application
- Targeting The Recessive Rice Gene, Xa13, For Bacterial Blight Resistance To A 14.8 Kb DNA Fragment
- Gene Expression In Giant Cells Induced In Tomato Roots By Meloidogyne Javanica
- Role Of Soybean E Genes In Regulating Onset And Duration Of Leaf Senescence
- Molecular Cloning And Characterization Of The TaLon1 In Wheat
- Isolation And Characterization Of A Gibberellin Responsive Gene (HvGR) From Initiated Shoots From Calli Derived From Mature Embryo Of Barley
- The Molecular Basis For Grain Texture In Wheat
- Can Transition To Flowering Be Modelled Dynamically From The Gene Level?
- Changes In Myo Inositol 1 Phosphate Synthase Gene Expression And Phytic Acid Accumulation In Oat Plants During Seed Maturation.
- Isolation Of Genes Involved In The Production Of Neurotoxin In Vetch, Vicia Sativa L.
- Proteomic Analysis Of Rice Caryopsis Development And Its Response To High Temperature
- Development Of Contiguous Introgression Lines Covering Entire Genome Of The Sequenced Japonica Rice
- Discovering Stay Green Drought Tolerance Genes In Sorghum: A Multidisciplinary Approach
- Genotype By Environment Interaction
- Environmental Effects On Phenotypic Expression Are Blunted In Greenhouse Compared To Open Field
- International Adaptation Trial: Using Probe And Reference Genotypes To Characterize Global Spring Wheat Production Environments.
- Interpreting QTL X Environment Interaction And QTL Specificity To Environments For Kernel Number In Winter Wheat With An Environmental Characterization Based On Probe Genotypes
- Measurement And Management Of Genotype Environment Interaction (GxE) For The Improvement Of Rainfed Lowland Rice Yield In Cambodia.
- 840_lofflercm
- Gene Discovery
- Genetic Resources For The 21st Century
- Monocots
- Identifying The Source Of New Variation Seen In Synthetic Backcross Derived Bread Wheat
- SSR Markers In Sampling A Core Collection And Estimating The Genetic Diversity
- Research On Genetic Diversity And Phylogeny Of Saccharum Spontaneum L. In China
- Genetic Diversity In Allelopathic Rice Accessions (Oryza Sativa L.)
- Identification And Mapping Of RAPD Markers Linked To Rice Stripe Virus Resistance Gene, Stvb I , In Rice(O. Sativa L.)
- Genetic Diversity Among Hexaploid Wheat Landraces With Different Geographical Origins Revealed By Microsatellites: Comparison With AFLP, And RAPD Data
- Collection And Conservation Of Genetic Resources For Dryland Farming Systems
- Using Of Wild Species Genetic Diversity In Plant Breeding.
- Diversification Of Australian Sorghum Using Wild Relatives
- Dicots
- Wild And Cultivated Cicer Species Different Evolutionary Paths Lead To Different Phenological Strategies That Can Be Exploited To Broaden The Adaptation Of Chickpea (C. Arietinum L.)
- Evaluation Of Genetic Diversity In Different Genotypes Of Brassica Juncea By SDS PAGE
- The Basis Of Sustainable Soybean Production: The Establishment And Application Of Chinese Soybean Core Collections Revealed By Both Agronomic Characters And SSR Markers
- Comparisons Of Growth And Photosynthetic Characteristics Between Wild And Cultivated Types Of Soybeans
- Use Of Primitive Accessions In Cotton Improvement
- Discovery Of New Diploid Perilla Species In Korea
- Genetic Diversity Of Nodules And Of Landraces Of Lupins Collected In Morocco Over The Last Five Years
- Homoclime Analysis Of Chickpea And Its Implications To Queensland’s Chickpea Industry
- Monocots
- Harnessing Molecular Markers For Plant Breeding
- Molecular Marker Selection
- Molecular Breeding For Rainfed Lowland Rice In The Mekong Region
- Diversity Arrays Technology, A Novel Tool For Harnessing Crop Genetic Diversity
- Mapping Genetic Loci Associated With Nitrogen Use Efficiency In Rice (Oryza Sativa L.)
- Combining High Protein Quality And Hard Endosperm Traits Through Phenotypic And Marker Assisted Selection In Maize
- QTL Analysis For Lodging Resistance In Rice Using A DH Population Under Lowland And Upland Cultural Conditions
- Molecular Markers For Selected Quality Traits In Australian Hexaploid Bread Wheat
- Tagging QTLs For Maximum Root Length In Rainfed Lowland Rice By Combined Selective Genotyping And STMs Markers
- Pedigree Based Genome Mapping For Marker Assisted Selection And Recurrent Parent Recovery In Wheat And Barley
- Molecular Breeding For Rainfed Lowland Rice In The Mekong Region
- QTL Identified For Yield Components In A Cross Between A Sugarcane (Saccharum Spp.) Cultivar Q165A And A S. Officinarum Clone IJ76 514
- Identification Of Genomic Regions Associated With Durable Stripe Rust Resistance In Wheat Line 11IBWSN50
- Marker Development And Implementation For Anthracnose Resistance In Australia Lupin Breeding Program
- The Asian Maize Biotechnology Network: Achievements And Opportunities
- Identification Of Quantitative Trait Loci For Flowering Time Using SSR Markers In Maize Under Water Stressed Conditions
- Molecular Characterization Of The S Haplotypes Lacking SLG In The Genome Of Brassica Campestris (syn. Rapa) L.
- Linkage Mapping Of QTLs For Seed Yield, Yield Components And Developmental Traits In Pea (Pisum Sativum L.)
- Mapping Of QTLs For Domestication Related And Agronomic Traits In A Temperate Japonica Weedy Rice
- Discovery Of SNPs In Soybean Genotypes Mainly Used As The Parents Of Mapping Populations In The USA And Korea
- Cross Incompatible Groups And Genetic Variation By RAPD Of Korean Sweetpotato [Ipomoea Batatas, (L.) Lam.] Varieties
- Development And Characterization Of Simple Sequence Repeat (SSR) Markers For An Apomictic Species Allium Senescens
- Identifying QTL Linked Markers In Marker Deficient Crops
- 802_basusm
- QTL Analysis For Grain Quality Properties In A Japonica Rice Combination
- Identification And Mapping Of RAPD Markers Linked To Rice Stripe Virus Resistance Gene, Stvb I , In Rice(O. Sativa L.)
- Markers Linked To A Grain Dormancy QTL In Wheat
- Tissue Culture
- Studies On Microspore Culture Of Hybrid Parents In Brassica Napus
- Effects Of Abscisic Acid On Callus Induction And Regeneration Of Different Wheat Cultivars To Mature Embryo Culture
- Development Of An Efficient High Frequency Microspore Embryo Induction And Doubled Haploid Generation System For Indian Mustard (Brassica Juncea)
- Embryogenesis From Isolated Microspores Of Chickpea And Field Pea – Progress Towards A Doubled Haploid Protocol As A Tool For Crop Improvement
- Using Tissue Culture To Select For Drought Tolerance In Bread Wheat
- Pollen Tube Behaviour And Effect Of Wheat Genotypes On Embryo Induction In Wheat X Maize Crosses
- Effect Of Physical, Chemical And Light Treatments On Germination And Growth Of Tissue Cultured Coconuts
- Effect Of Genotype On Adventitious Bud Formation From Leaves Of Rapeseed (Brassica Napus L.)
- Influence Of Age Of The Embryo And Method Of Hormone Application On Haploid Embryo Formation In Wheat X Maize Crosses
- Utilization Of Anther Culture Technique For Rice Improvement In The Philippines
- Tanbaguro: A New Model Genotype Of Soybean For Tissue Culture Study
- Effect Of Plant Growth Regulators On Direct Shoot Regeneration Of Wheat (Triticum Aestivum)
- Interspecific Crossing
- Cytomorphological Studies In Genus Citrullus (Cucurbitaceae)
- Chromosome Substitution Lines From Gossypium Barbadense L. As Sources For G. Hirsutum L. Improvement
- Cross Compatibility Of Elite Papaya Inbred Lines To An Intergeneric Hybrid Of Carica Papaya L. X Vasconcella Quercifolia (Saint Hil.) Hieron
- Male And Female Gamete Abortions, And Reduced Affinity Between The Uniting Gametes As The Causes For Sterility In An Indica/japonica Hybrid In Rice
- Development Of Lowland Rice From The Interspecific Cross Of Oryza Sativa And O. Glaberrima
- Heterosis Rice And Other Crops
- Gene Pyramiding To Improve Hybrid Rice By Molecular Marker Techniques
- Critical Temperature And Stages Of Fertility Alteration In Thermo Sensitive Genic Male Sterile Lines Of Rice
- Investigation Of Maize Heterotic Groups And Patterns In China
- Exploitation Of Heterosis For Raising Productivity In Sesame
- ISSRs: An Efficient Tool To Characterize Interspecific F1 Hybrids Of Brassica Species
- Testing Validity Of Fertility Restorer (Rf ) Gene Associated RAPD Markers In Newly Identified Restorer And Maintainer Lines Of Pepper (Capsicum Annuum L.)
- Cloning And Characterisation Of Rf1, A Fertility Restorer Gene For Ms Bo Type CMS Rice
- Load Of Deleterious Genes In Maize Estimated By Inbreeding Depression, Yield Potential Per Plant And CV
- Induction Of Male Sterility In Niger (Guizotia Abyssinica Cass.)
- RAPD And Isozyme Markers For Genetic Diversity And Their Correlation With Heterosis In Rice (Oryza Sativa L)
- Simulation Of Positive Assortative Mating For Inbred Line Development Using QU GENE
- Genetic Analysis
- Combining Ability For Some Quantitative Characters In Hexaploid Wheat (Triticum Aestivum L. Em. Thell)
- Inheritance Of Biochemical Characteristics In Non Premature Senescing, Short Seasoned Cotton
- Inheritance Of Seed Isoflavone Contents In Soybean Cultivars Shinpaldalkong2 And Hwangeumkong
- Inheritance Of Kernel Elongation In Rice
- Genetic Analysis Of Seedling Characters In Bread Wheat
- Genetic Analysis Of Plant Height And Panicle Erectness Of Rice
- Secondary Dormancy Of Oilseed Rape: First Aspects Of Heredity
- Efficient Evaluation Of Lines Derived From Recurrent Selection In Barley
- Yearly Variation Of Genetic Parameters For Panicle Characters Of Japonica Rice (Oryza Sativa L.)
- Novel Breeding Approach Through Development Of Introgression Lines In Rice
- Strategies For Breeding Against Barley Grain Colour Defects
- Combining Ability Of CIMMYT’s Early Maturing Maize (Zea Mays L.) Germplasm Under Stress And Non Stress Conditions And Identification Of Testers
- Estimate Of Some Genetic Parameters In Corn (Zea Mays L.) Based On Diallel Crossing System
- Physiological Traits
- Genetic Analysis Of Phenological Traits And Grain Yield In Durum Wheat Under Normal And Late Sown Environments
- Can Canopy Temperature Depression Measurements Help Breeders In Selecting For Yield In Wheat Under Irrigated Production Conditions?
- Seed Coat Integrity: Inheritance And Selection For Resistance To Mechanical Damage In Common Beans (Phaseolus Vulgaris)
- Integrating Crop Physiology Into The Australian Soybean Improvement Program
- Selection Of Parents For Studying B Tolerance In Brassica Rapa
- Variety Identification, Release, Progress
- Is There A Case For Discarding Low Yielding Soybean Performance Trials?
- Study On Rice Breeding Process For High Yielding And Good Quality Varieties
- Recent Advances In Crop Improvement In China
- Seeds Of Life Increasing Production Of Staple Crops In East Timor
- Lentil Improvement For The Benefit Of Highland Farmers
- Development Of Early Maturing Chickpea Varieties For Diversification Of Rice Wheat Cropping System
- High Protein Lupins: Diversifying The Pulse Industry In Western Canada.
- Early Maturing Sweet Potato Varieties For High Altitude Highlands Of Papua New Guinea
- Breeding For Increasing Wheat Yield In The Cold Dryland Regions Of Iran
- Using Corn Hybrid Yield Data To Improve Selection Of Rapidly Changing Hybrids
- An Effective Procedure To Maintain A Bread Wheat Cultivar
- Molecular Marker Selection
- New Frontiers And Products From Breeding
- Superior Corn Based Starches For Oil Field Application
- Effects Of Planting Time And Plant Density On Flower Yield And Active Substance Of Chamomile (Matricaria Chamomilla L.)
- Jute Quality Breeding By Reducing Lignin Content
- Introgression Of Desaturase Suppressor Gene(s) From Brassica Napus L. To Enhance Oleic Acid Content In Brassica Juncea L. Coss.
- 596_hoppnerf
- New Lines Raise The Commercial Potential Of Guayule
- Abiotic Challenges And Breeding
- General
- Salt
- Recovery Strategy Of Plant Growth And Yield Following The Foliar K And Ca Applications On The Salt Stressed Rice Plant
- 1147_boughanmi
- Screening Chickpea (Cicer Arietinum L.) And Wild Relatives Germplasm From Diverse Country Sources For Salt Tolerance
- Differential Response Of Wheat Cultivars To Subsoil Salinity/sodicity
- Subsoil Salts Affect Root Function, Shoot Growth And Ionic Balance Of Wheat Plants
- Expression Of Ectoine Biosynthetic Genes In Tobacco Plants (Nicotiana Tabaccum) Leads To The Maintenance Of Osmotic Potential Under Salt Stress
- Effects Of NaCl Salinity On Growth And Bt Protein Content Of Two Transgenic Bacillus Thuringiesis Cotton Cultivars
- Light Dependency Of Salinity Induced Chloroplast Damages
- Salt Stress Magnitude Can Be Quantified By Integrating Salinity With Respect To Stress Duration
- Oxygation Of Rhizosphere With Subsurface Aerated Irrigation Water Improves Lint Yield And Performance Of Cotton On Saline Heavy Clay Soil
- Salinity Induced Chloroplast Damages In Rice Leaves (Oryza Sativa L.) Are Reduced By Pretreatment With Methyl Viologen
- Validation And Application Of Salt Stress Unit In Dealing With Salt Affected Growth And Leaf Proline Accumulation Of Rice Plants
- Temperature
- Immunological Assays Of Freezing Tolerance In Barley Using Antifreeze Proteins Antisera
- Monitoring Characteristics Of Cold Hardiness For Perennial Ryegrass Organs From The View Point Of Dynamic States Of Water
- Studies On Boundary Of Winter Wheat Northernmost Planting In Chinese Northeast Area
- Effect Of Delayed Planting On Membrane Injury And Yield Of Six Chickpea Genotypes
- Investigation Of Post Head Emergence Frost Resistance In Several CIMMYT Synthetic And Queensland Wheats
- Expression Of Genotypic Responses To Frost Tolerance In Wheat
- Regulation Of Freezing Resistance In Barley Grown Under Field Conditions
- Correlation Between Reliability Of Pollination And Length Of Basal Dehiscence Of The Thecae In Rice Under A Hot And Humid Condition
- Other: Heavy Metal, Waterlogging, Drought
- Comparison Of Root Staining And Root Elongation In Predicting Aluminium Tolerance Using SSR Markers In Barley
- Identification Of Physiological Indicators For Establishing Screening Techniques Related To Tolerance Of Excess Water In Soybean [Glycine Max(L.) Merrill].
- Accumulation Of Ammonium Ion In Cadmium Tolerant And Sensitive Cultivars Of Oryza Sativa
- Kernel Set In A Maize Hybrid And Its Two Parental Inbred Lines Exposed To Water Stress.
- Inheritance Of Waterlogging Tolerance Of Barley (Hordeum Vulgare L.)
- Grain Size Distribution Is Related To Small Grain Screenings In Wheat
- Genetic Evaluation And Improvement Of Acid Stress Tolerance In Lucerne Breeding
- Assessing The Impact Of High Soil Boron And Salinity On The Root Growth Of Potential Primer Plant Species
- Effects Of Foliar Fertilizer Applications On The Alleviation Of Waterlogging Stress In Six Barley Varieties
- Responses To Boron (B) Levels In F2 And F3 Populations Derived From B Efficient And B Inefficient Wheat Parents
- The Iron (Fe) Excluding Power Of Rice Roots As A Mechanism Of Tolerance Of Elite Breeding Lines To Iron Toxicity.
- Biotic Challenges And Breeding (5ACSC)
- Insects, Nematodes
- Saponins In Alfalfa And Their Relationships With Alfalfa Weevil Resistance
- Influence Of Seed Origin On Scale Insect (Maconellicoccus Hirsutus) Damage And Severity In Teak Provenance Trial
- Influence Of Cytoplasmic Male Sterility On The Expression Of Resistance To Insects In Sorghum
- Genes For Resistance To Russian Wheat Aphid In Two New Wheat Lines
- Progress In Genetics And Mapping For Resistance In Soybean [Glycine Max (L.) Merrill] To Cyst Nematode (Heterodera Glycines Ichinohe).
- Fungii, Monocots
- Sorghum Ergot: Revealing The Genetic Architecture Of Resistance
- Development Of International Standard Differential Set For Blast Resistance In Rice (Oryza Sativa L.)
- Strategy For The Identification And Breeding Of Resistance To Dryland Root Rot Complex For International Spring And Winter Wheat Breeding Programs.
- Dryland Root Rot: A Major Threat To Winter Cereal Production Under Sub Optimal Growing Conditions
- Relationships Between Resistance To Fusarium Head Blight And Crown Rot In Hexaploid Wheat
- Fungii Dicots
- International Collaboration To Develop Integrated Management And Resistant Varieties To Minimise The Impact Of Botrytis Grey Mould In Chickpea
- Introgression And Biochemical Manifestation Of The Gene(s) For White Rust Resistance In Indian Mustard (Brassica Juncea (L.) Coss.)
- Discovering And Characterising The Genetic Mechanism Controlling The Resistance Response To Ascochyta Blight (Ascochyta Rabiei) In Chickpea (Cicer Arietinum)
- New Ascochyta Blight Resistant, High Quality Kabuli Chickpea Varieties For Australia
- Identification Of RGAs From A Pea BAC Library Using BAC Pools
- Dunkeld And Rainbow: A Tale Of Two Victorian Canola Cultivars
- Viruses, Bacteria
- QTLs Associated With Resistance To The Cassava Mosaic Disease
- Misunderstandings On Parasitic Weed Control And Research In The World With The References Of Striga Tolerance In Maize
- Genetic Analysis Of Resistance To The Cassava Mosaic Virus Disease
- Gene Expression Of Lactoferrin Derived Antimicrobial Peptides In Rice
- Evaluation Of Bacterial Blight Resistance In Rice Lines Carrying Multiple Resistance Genes And Xa21 Transgenic Lines
- Molecular Mapping Of Resistance To Sugarcane Mosaic Virus In Maize
- Development Of Japonica Rice “Iksan478” Carrying Xa7 Resistant Gene To Bacterial Blight
- Towards Development Of PRSV Resistant Papaya By Genetic Engineering
- Insects, Nematodes
- Biotechnology For A Better World
- Production And Characterization Of Recombinant Human Lactoferrin In Transgenic Javanica Rice Cv Rojolele
- Grain Growers’ Perspectives On Genetically Modified Organisms (GMOs)
- Gene Transformation In To Dicotyledonous Plants By Using Agrobacterium And Study On Its Problem
- Harnessing RNA Silencing To Protect Peanuts From Stripe Disease
- Studies Of Differential Sensitivities Between Conventional And Transgenic Bt Cotton Cultivars To Potassium Deficiency
- Effects Of Nitrogen Fertility On Bt Endotoxin Levels In Bt Hybrid Maize
- Transformation Of Vigna Mungo (blackgram) For Abiotic Stress Tolerance Using Marker Free Approach
- Molecular Gene Transfer For The Generation Of Salt Tolerant Rapeseed (Brassica Campestris, Brassica Napus) Varieties In Bangladesh
- Genetic Transformation Of Russian Wheat Varieties For The Biotic And Abiotic Stress Resistance
- Effectiveness Of Four Cry Genes In Transgenic Potato For Conferring Resistance To Potato Tuber Moth
- Public Perceptions About Genetically Engineered Forage Crops And Resultant Animal Products
- The Development Of Virus Resistant Carrot Genotypes Using RNAi Technology
- Glyphosate Alters Floral Morphology In Glyphosate Resistant Cotton
- Genetically Engineered Tomatoes: New Vista For Sustainable Agriculture In High Altitude Aegions
- Genome Sequencing
- 4. Effecting Change
- Proven Processes For Change
- Developed Countries
- Ricecheck –Participatory Farmer Extension Model In Practice For 18 Years
- Optimising Profitability And Environmental Trade Offs In Managing Cropping Systems Using Differential Evolution
- Decision Support Systems For Farm Management: A Theoretical Framework From The Sociology Of Science And Technology.
- Farmer Led On Farm Trials – Problem Or Solution?
- State Focus – Improving Farmer Practices Through Coordinated On Farm Testing.
- National WhopperCropper – Risk Management Discussion Support Software
- Computer Based Decision Support Tools For Australian Farmers
- Agro Biotechnology Outreach Program Using SACUC Model
- Best Management Practices – A Multi Disciplinary Approach To Improving The Yield Of High Input Peanut Production In Australia
- Developing Countries
- 1233_maboudou
- Peanut Farmers’ Experience With FARMSCAPE In Peninsular India
- Technology Dissemination To Boost Pulse Production And Human Nutrition In Bangladesh
- Participatory Varietal Selection (PVS) To Identify Upland Rice Cvs In Ghana
- Participatory Research For Sustainable Yield Improvement Of Rice In Rainfed Lowlands: A Case Study Of Eastern U.P., Eastern India
- An Example Of Participatory Plant Breeding: Barley At ICARDA
- Farmer Participation In Groundnut Rosette Resistant Varietal Selection In GHANA
- 664_affholderf
- Diagnosing Variable Nutrient Deficiencies In Rainfed Lowland Rice Using Strip Trials
- Developed Countries
- Lessons For Future From Success Stories
- Role Of IT In Delivery And Adoption
- Spatial And Temporal Stability Of Corn Grain Yields In The Upper Rhine Valley
- Development Of A Matrix Based, Multi Media Diagnostic Key To Sweetpotato Problems
- Analysis Of Spatial Variation Of Rice Grain Yield And Soil Chemical Properties
- Gathering And Sharing Crop Science Knowledge In A Global Community Of Practice
- New Pathways For Delivery In Asia 5ACSC
- Planning, Monitoring And Evaluation Of Research
- Private/public Complementation
- Proven Processes For Change
- 5. Asian Crop Science Society
- Food Quality Issues
- Rice
- New Rice Cultivars With Low Levels Of Easy To Digest Protein
- Nitrogen Fertilizer Alters Milling Quality And Protein Distribution In Head Rice
- Development Of Special Purpose Aromatic Rice Varieties In The United States
- Grain Quality And Iron Density Of Philippine Rice Cultivars
- Variation In Grain Iron Between Seed Lots Of Some Upland Rice Varieties
- Stability Of Tocopherols And Tocotrienols Extracted From Unsaponifiable Fraction Of Rice Bran Under Various Temperature And Oxygen Condition
- Early Identification Of Pre Harvest Sprouting In Rice Seeds By Using 1H NMR
- Chemical And Sensory Evaluation Of Aromatic Rice Cultivars.
- Natural Variation In Alk Locus Affects Starch Property And Cooked Rice Quality
- Changes Of Anthocyanin Pigment Cyanidin 3 Glucoside, Oryzanol Content And Antioxidant Activity As Affected By Ripening Temperature In Rice Varieties
- Maize, Wheat
- Prospects Of Breeding Sorghum, Pearl Millet And Pigeonpea For High Forage Yield And Quality
- Variety And Environment Effects On Quality Traits In Latvian Grown Winter Wheat
- Effect Of Fertilizer Treatment On The Content Of Protein Composition And The Processing Quality Of Bread Wheat
- The Effects Of Elevated Temperature On Starch Deposition In Developing Wheat Grains
- Selecting Maize Varieties For Eating Quality In East Timor
- The Effects Of Elevated Temperature On Starch Deposition In Developing Wheat Grains
- 617_srinivasang
- Other
- Canola Quality Brassica Juncea For Australia.
- Making A Greener Revolution: A Nutrient Delivery System For Food Production To Address Malnutrition Through Crop Science
- Associations Between Isoflavones And Protein Content In Soybean (Glycine Max) Seed
- Community Based Fish Culture In Seasonally Flooded Rice Fields In Bangladesh And Vietnam
- Investigation Of Biological Effect In Black Colored Soybean
- Selection Of Sweetpotato Clones With High ββ Carotene Content
- Aflatoxins And Their Relationship With Sugars In Peanut (Arachis Hypogaea L.)
- Diminution Effect On Mycotoxin Content By Different Processing Methods In Barley And Wheat Infected With Fusarium Graminearum
- Assessment Of The Impact Of Arsenic Containing Irrigation Water On Soil Contamination And Plant Uptake
- A New Soybean Cultivar, “Fukuibuki”, With High Isoflavone Content And Superior Agronomic Characteristics For Japan
- Development Of Nutritionally Superior Brassica Napus And B. Juncea Oils Using RNAi Mediated Gene Silencing
- 813_ruhl
- Rice
- Feed Quantity And Quality Issues
- Prospects Of Breeding Sorghum, Pearl Millet And Pigeonpea For High Forage Yield And Quality
- Economic Sustainability: Genetic And Cutting Schedule Influences On The Yield Quality Tradeoff In Alfalfa (Medicago Sativa L)
- Effects Of Different Cutting Stages On Forage Yield And Quality Of Nilegrass And Pangola Grass1
- Spring Regrowth Of Lucerne Is Affected By The Level Of Winter Perennial Reserves.
- Comparison Of Digestibility And Metabolizable Energy Between Nilegrass And Pangola Grass
- Desmanthus – A New Pasture Legume For The Dry Tropics
- Plant Traits Associated With Winter Growth Of Perennial Grass Cultivars
- Shrivelled Grain Potentially Worth More Than Chicken Feed
- Cutting Management Of Multipurpose Tree Legumes: Effects On Green Herbage Production, Leaf Retention And Water Use Efficiency During The Dry Season In Timor, Indonesia
- Performance Indicators For Harvest Timing Of Whole Crop Cereals For Silage
- Food Quality Issues
- Symposia Papers
- One
- Breeding Crops That Do Better Under Water Scarcity
- Irrigated Rice Based Systems 5ACSC
- Physiology And Improving Productivity Under Water Scarcity
- Dryland Cropping Systems
- Water Harvesting And Supplemental Irrigation For Improved Water Productivity Of Dry Farming Systems In West Asia And North Africa
- Management Of Extensive Farming Systems For Drought Prone Environments In North America And Australia
- Economics Of Rainwater Harvesting For Crop Enterprises In Semi Arid Areas
- New Imperatives For Irrigated Agriculture
- Water And Nutrients In Turf Grass
- Environmental Consequences Of Water Management In Farming Systems
- 2. Crop Science For A Sustainable Future
- Farming And Cropping Systems
- Conservation And Reduced Tillage Systems
- Turfgrass, The Most Visible Crop
- Asian Cropping Systems (5ACSC)
- Biocide Dependence And Resistance
- Nutrient Recycling And Balance In Cropping Systems
- Progress In Lifting Soil Fertility In Southern Africa The Views Expressed In This Paper Are Those Of The Authors’ And Do Not Necessarily Reflect Those Of CIMMYT Or DARS. The Information Summarized Here Has Come From The Work Of Many Members Of Soi
- Nutrients In Organic Farming – Are There Advantages From The Exclusive Use Of Organic Manures And Untreated Minerals?
- Managing Residues And Nitrogen In Intensive Cropping Systems. New Understanding For Efficient Recovery By Crops.
- Managing Climate Risk In Cropping Systems
- Raising Yield Potential In Cropping Systems
- 3. Crop Science For Harnessing Genetics
- Genome Sequencing Of Model Systems
- Gene To Phenotype Complexity
- Equation Chapter 1 Section 1Complex Trait Genetics And Gene To Phenotype Models
- Equation Chapter 1 Section 1Modelling Gene Networks Controlling Transition To Flowering In Arabidopsis
- Trait Physiology And Crop Modelling To Link Phenotypic Complexity To Underlying Genetic Systems.
- Modelling Differential Phenotypic Expression
- The Struggle To Exploit Non Additive Variation
- 778_tardieuf
- Genetic Resources For The 21st Century
- Harnessing Molecular Markers For Plant Breeding
- Use Of Marker Assisted Selection In A Product Development Breeding Program
- Applications Of Genomics Technologies To Enhance Rate Of Genetic Progress For Yield Of Maize Within A Commercial Breeding Program
- Implementation Of Molecular Markers For Quantitative Traits In Breeding Programs Challenges And Opportunities
- New Frontiers In Altering Plant Metabolism
- Adaptation To Abiotic Challenges
- Adaptation To Biotic Challenges (5ACSC)
- Biotechnology For A Better World
- 4. Crop Science For Effecting Change
- Proven Processes For Innovation And Change
- The Evolution Of Extension Processes And Practices In Relation To Smallholder Farming In Southern Africa
- Participatory Research: Does It Work? Evidence From Participatory Plant Breeding
- Decision Support Systems In Australian Dryland Farming: A Promising Past, A Disappointing Present And Uncertain Future
- Implementing Change Lessons For The Future
- The Role Of IT In Enhanced Delivery And Adoption
- New Pathways For Delivery In Asian Countries (5ACSC)
- Research Planning Monitoring And Evaluation
- Agents For Change Public, Private And NGOs
- Proven Processes For Innovation And Change
- 5. 5th Asian Crop Science Conference
- 6. 12th Australian Agronomy Conference
- Pathways To Profitable Agriculture
- Removing Soil Compaction And Increasing Water Use Efficiency.
- The Changing Face Of Agriculture In Australia
- New Ascochyta Blight Resistant, High Quality Kabuli Chickpea Varieties For Australia
- A Long Term View Of Australian Agriculture: History, Future And Lessons
- Simulating Lucerne/crop Companion Farming Systems In Australia
- Farming And Land Stewardship
- Boron Tolerance Of Lentil Highlights Of A Research Program
- Fallow Management Affects The Risk Of Deep Water Loss
- Simulation Of Nitrogen Management In Trash Blanketed Sugarcane Systems
- Farming And Land Stewardship. Case Study – Australia’s Innovations In Sustainable Irrigation
- Risk Assessment Of Climate Change Impacts On Australia’s Wheat Industry
- Crop Livestock Interaction
- Grain Graze As Sustainable Farming Systems In Sub Tropical Queensland
- The Benefits And Challenges Of Crop Livestock Integration In Australian Agriculture
- Potential Of Some Annual Pasture Legumes As Fodder Crops In The Wheatbelt Of Western Australia
- The Whole Farm Impact Of Including Dual Purpose Winter Wheat And Forage Brassica Crops In A Grazing System: A Simulation Analysis
- Genetic Improvement Of Strand Medic (Medicago Littoralis Rohde Ex Lois.) For Australian Farming Systems
- Research Implementation And Delivery
- Pathways To Profitable Agriculture
- One
- 2006 13th AAC
- Concurrent Papers
- Participatory Research: Adopting Agronomy
- Evaluating Knowledge And Practices To Combat Subsoil Constraints In North Eastern Australia
- Developing A Better Understanding Of Determining Potassium Requirements In Cereal Cropping In The Central Wheatbelt Of Western Australia.
- Participatory Research For Agronomy: The New Zealand Experience
- Factors Affecting Planting Decisions In Central Queensland Farming Systems Lessons From On Farm Research
- 4645_huntj
- 4658_williams
- 4660_bestf
- Lifetime Wool: Developing Effective Extension Packages For Farmers A Case Study Based On Sustained Behaviour Change Principles
- Improving Crops And Pastures For Targeted Environments
- Plant Parameters Identified For Evaluating Varietal Performance Of Acid Tolerance In Lucerne
- Development And Adoption Of New Kabuli Chickpea Varieties In Australia
- On Wheat And Salmon: The Trade Off Between Seed Size And Number
- Poor Persistence Of Sub Tropical Grasses Over Winter
- Changes In Sorghum Yield Components After Chilling
- Improving Forage Legume Options For Saline Environments – Melilotus Species
- Tapping The Large Genetic Variability For Salinity Tolerance In Chickpea
- Screening For Sodium Exclusion In Wheat And Barley
- Reduced Tillering Wheat Lines Maintain Kernel Weight In Dry Environments
- Reducing Management Inputs And Maximising Seed Quality In Faba Beans Through Improved Varieties
- Screening For Waterlogging Tolerance Of Wheat In The Field In Western Australia
- Hardy Australians: Ecogeography Of Cullen Suggests Perennial Legumes For Low Rainfall Pastures
- Agronomy As Applied Ecology Or Why We Shouldn’t Lose Sight Of The Big Picture When Marking The White Pegs. A Chickpea Example.
- Overcoming Soil And Nutrient Constraints
- Hardpan Penetration Ability Of Wheat Roots
- Root Turnover And Microbial Biomass In Cotton Farming Systems
- Understanding Deep Drainage In Grey Vertosols Of The Moonie And Condamine Catchments Of South East Queensland
- High Chloride In Subsoil: A Key Indicator For Potential Grain Yield Losses In Southwest Queensland
- 4555_daviess
- Towards An Understanding Of Variability In Yield Responses To Liming
- Catchment Based Long Term Monitoring Reveals The Value Of Lime Application To Correct Soil Acidity.
- Rooting Patterns In Double Haploids Lines Of Wheat For Reduced Tillering And Their Relationship With Early N Uptake
- Identifying Nutrient Management Opportunities And Threats In The Western Australian Grains Industry
- Response Of Wheat To Split Application Of Nitrogen On A Leaching Sandy Soil
- Subsoil Constraints And Wheat Production In Queensland: Economic Impacts And Fertiliser Management Opportunities
- Are Flip Flop Yields Of Crops On Alkaline Soils In The Victorian Mallee Related To Subsoil Physicochemical Constraints?
- Simulating The Impacts Subsoil Constraints On Wheat Yields In The Northern Grain Zone
- The Effect Of P And K Nutrition On The Tolerance Of Wheat And Chickpea To Subsoil Salinity.
- Building Crop And Pasture Systems For Sustained Productivity And Profitability
- Short Term Cover Crops Increase Wheat Yields In Southern Queensland.
- Competitive Effects Of Wild Radish (Raphanus Raphanistrum L.) On Lupin Cultivars (Lupinus Angustifolius L.)
- Who Benefits Most From ‘public’ Herbicide Use R&D?
- Hairy Canary Clover: A Case Study For Integrating Semi Herbaceous Forage Plants Into Agricultural Systems
- 4511_parsonsd
- Management Of The Interface Between Sugarcane Cycles In A Permanent Bed, Controlled Traffic Farming System.
- Using The Weed Seed Wizard To Understand And Manage The Weed Seedbank
- Manipulating Row Spacing To Improve Yield Reliability Of Grain Sorghum In Central Queensland
- Fenugreek Has A Role In South Eastern Australian Farming Systems
- Loss Of Bioavailable Pendimethalin Herbicide From Soil Under Different Seeding Systems
- Differences Among Wheat Cultivars In Their Optimum Sowing Times In Western Australian Environments
- A Role For Wide Rows In Lupin Cultivation In Western Australia
- Turning Pastures Into Profit: Optimising Pasture Utilisation And Stocking Rates
- Do Ultra Narrow Row Cotton Systems Offer Any Benefits To Australian Farmers?
- 4571_whitepf
- 4588_akponikpep
- Assessing The Feasibility Of GM Cotton In The Ord River Irrigation Area: Tillage Systems For Late Wet Season Sowing.
- Developing Supplementary Feed Systems For Dairying In Dry Environments In New Zealand
- Managing High Stubble Loads: Is Grazing The Answer?
- Data Mining The WhopperCropper Database: Sorghum In Central Queensland.
- Modelling The Population Dependence Of Canopy Development And Biomass Partitioning Of Field Peas (Pisum Sativum L.)
- Effect Of Butterfly Pea Ley Pastures On Subsequent Wheat Crops
- Plant Diversity In Vegetable Cropping Systems
- Economic Chickpea Production For Southern Australia Through Improved Cultivars And Strategic Management To Control Ascochyta Blight.
- Adapting Crop Species On Grey Vertosols In North West NSW To Compensate For High Chloride Levels In The Subsoil.
- Dual Purpose Canola – A New Opportunity In Mixed Farming Systems?
- Assessing The Impact Of Agronomy On Wheat Yield Across Soils And Locations
- Wheat Yields After Perennial Pastures Under Drought Conditions
- Managing Spatial Variability Of Grain Yield At The Paddock Level In Southeastern Australia
- Interpreting Variable Data From Herbicide Tolerance Trials On Pulse Cultivars For On Farm Applicability.
- The Future Of No Tillage Systems In Western Australia
- Dual Purpose Long Season Winter Wheats To Improve Productivity In Western Australia
- A Diagnostic Approach To Cropping Systems Research: Two Sites At South Stirling, WA.
- Impact Of Row Configuration On High Fruit Retention (transgenic) Rain Fed Cotton Systems
- Future Technologies In Agriculture
- Technologies To Estimate Plant Available Soil Water Storage Capacities At High Spatial Resolution
- 4592_carrs
- Advances And Challenges In The Spatial Detection Of Nitrogen And Water Stress In Wheat
- Oxygation: Enhanced Root Function, Yields And Water Use Efficiencies Through Aerated Subsurface Drip Irrigation, With A Focus On Cotton.
- Managing Water In A Changing Environment
- Soil Water Under Lucerne Phase And Companion Cropping Systems.
- Reduced Frost Damage In Wheat Crops Grown In Delved Soils
- An Alternative Method For Simulating Crop Water Use
- Comparative Water Use Productivity Of Forages For The Dairy Industry In Northern Victoria
- PAWC Determines Spatial Variability In Grain Yield And Nitrogen Requirement By Interacting With Rainfall On Northern WA Sandplain
- Water Extraction Of High Retention Cotton Crops
- Application Of APSIM ‘multi Paddock’ To Estimate Whole Of Farm Water Use Efficiency, System Water Balance And Crop Production For A Rice Based Operation In The Coleambally Irrigation District, NSW
- Initial Soil Water: Myth And Management
- Options For Root Zone Drainage In High Rainfall Areas.
- The Influence Of Grazing Interval And Perennial Grass Species On Soil Moisture.
- On Farm Water Management And The Madden Julian Oscillation
- Participatory Research: Adopting Agronomy
- Conference Information
- Plenary Papers
- Participatory Research: Adopting Agronomy
- Improving Crops And Pastures For Targeted Environments
- Using The ROOTMAP Model Of Crop Root Growth To Investigate Root Soil Interactions
- Implications Of ‘duty Of Care’ For The Development Of New Pasture Species
- Genetic Diversity In Australian Canola And Implications For Future Breeding
- Invited Paper: Lupin Genetic Improvement For Targeted Environments And Markets
- Invited Paper: Using Synthetic Wheats To Breed Cultivars Better Adapted To Changing Production Conditions
- Opening Oration
- Overcoming Soil And Nutrient Constraints
- Building Crop And Pasture Systems For Sustained Productivity And Profitability
- New Annual Pasture Legumes For Southern Australia 15 Years Of Revolution
- The Role Of Nitrogen And In Crop Lucerne Suppression For Increasing Cereal Performance In Companion Cropping Systems
- Burning Narrow Windrows For Weed Seed Destruction
- Invited Paper: The Tension Between Specialization And Diversification In The Evolution Of Farming Systems For The Australian Crop Livestock Zone.
- Future Technologies In Agriculture
- Managing Spatial And Seasonal Variability Within Field Can Improve The Profitability Of WA Grain Production
- Remote Sensing To Detect Nitrogen And Water Stress In Wheat
- LUCI In The Sky With Diamonds – Modelling The Wider Impacts Of Land Use Change And Intensification
- Precision Agriculture In The Victorian Wimmera – Grower Perspectives
- Invited Paper: Appropriate Technologies For Agricultural Sustainability In An Uncertain Future
- Managing Water In A Changing Environment
- Poster Papers
- Participatory Research: Adopting Agronomy
- Participatory Evaluation By Farmers Of On Farm Seed Priming In Wheat In Assam, India
- How A Participatory Research Approach Has Improved The Viability Of Lupin Crops In Western Australia.
- Delivering Soil Water Information To Growers And Consultants
- Field Performance Of New Wheat Varieties In Large Scale Trials On The South Coast Of Western Australia
- Agronomic Performance And Adoption Of Liquid Fertilisers In Western Australia
- Value Adding To The Agribusiness Trial Network In Western Australia
- Local Delivery For Statewide Impact: Lessons From A Grower Group Alliance
- From Adaptation To Adoption – A Case Study Of Burgundy Bean
- Conservatism, Complexity And Consequences
- Integrated Crop Management Of Chickpea In Environments Of Bangladesh Prone To Botrytis Grey Mould
- Participation For Improved Adoption, Research, Or Both: Two Case Studies
- Modern Maize Varieties Are More Weevil Susceptible Than Local Populations When Stored In Local Manner In East Timor.
- Building Crop And Pasture Systems For Sustained Productivity And Profitability Agronomy
- Alternative Perennial Legumes For The High Rainfall Zone Of SE Australia.
- Effect Of Grain Size On The Competitive Ability Of Wheat
- Carrot Yield On A Steep Slope Versus Low Slope From The North West Coast Of Tasmania: A Preliminary Study.
- Preliminary Studies Of Inheritance Of Autogamy In Lotus Corniculatus L.
- Crop Characteristics For High Yield Of Wheat In The High Rainfall Zone Of Western Australia
- Seasonal Changes In Root Dry Matter And N Content Of Lucerne Crops Under Contrasting Defoliation Regimes
- Lucerne Canopy Expansion In Spring Was Affected By The Level Of Winter Root Reserves.
- Row Spacing Impacts On Dryland Wheat Yields In Central Queensland.
- Potential Of Tropical Forages For The Tasmanian Dairy Industry
- Agronomic Performance And Adoption Of Liquid Fertilisers In Western Australia
- Plant Characteristics Associated With Wheat Yield In The High Rainfall Zone Of Southern Australia
- Components Of Long Term Nitrogen Balance In Central Qld Grain Cropping Systems.
- Salt Priming Improves Establishment Of Distichlis Spicata Under Saline Conditions
- Evaluation Of Oat Varieties For Hay Production In Western Australia
- A Nitrogen Fertiliser Calculator Emphasising Climate Risk Management
- High Wheat Seeding Rates And Plant Density Do Not Go Hand In Hand
- Mycorrhizae: A Benefit To High Yield Maize Cropping Systems?
- Performance Of Two Wheats With Differing Row Configurations, And Nitrogen And Sowing Rates
- Impact Of A History Of Lucerne (Medicago Sativa L.) On The Root Dynamics Of A Subsequent Canola (Brassica Napus L.) And Wheat (Triticum Aestivum L.) Crop
- Increasing Fodder Production In Central Tibet
- Quantifying Carbon Sequestration On Saltland Pastures In South West Australia
- Agronomic Performance Of Dwarf Milling Oats In Western Australian Environments
- When To Sow Chickpea In South Western Australia
- Eight Years Of Distichlis Spicata Growth Improves Soil Properties In A Saline Discharge Zone
- Nitrogen Management For Increased Wheat Production In The High Rainfall Cropping Zone Of Western Australia
- Improving Crops And Pastures For Targeted Environments
- Trifolium Dasyurum L. (Eastern Star Clover): Exploiting A Novel Pasture Species For Innovative Strategies To Control Weed Seed Banks.
- Can Grain Protein Contents Be Increased Without Sacrificing Grain Yields In Wheat?
- Perennial Herbs – Can They Make A Contribution In High Rainfall Dairy Pastures?
- The Effect Of Dairy Effluent On Dry Matter Yields And Nutritive Characteristics Of Summer Active Regrowth Forage Crops
- Waterlogging Tolerance Of 12 Perennial Forage Legumes From The Lotus Genus
- Austrodanthonia Caespitosa Shows Potential As A New Pasture Grass For The Low Rainfall Wheatbelt
- Predicting Mixed Stand Alfalfa Fibre Content In New York State
- Production And Persistence Of A Winter Active Chicory Cultivar In Southern NSW
- Summer Dormancy Increases Persistence Of Perennial Grasses In A Wheatbelt Environment
- Deep Incorporation Of Organic Matter Improves Wheat Growth On A Sodosol With Dense Subsoil
- Melilotus Siculus (syn Messanensis) Is Constrained By A Lack Of Suitable Rhizobia
- Interspecific Hybrid, Zea Mays L. X Tripsacum Dactyloides L., A New Fodder Crop With Large Silage Biomass Production Under Abiotic Stresses
- Understanding Productivity Decline In Subterranean Clover Based Pastures In South Western Australia
- Quantifying Safflower Development Using Environmental Parameters
- Matching Agronomic Practices With Improved Cultivars To Expand Lentil Production In Australia
- Building Crop And Pasture Systems For Sustained Productivity And Profitability Pests, Diseases & Grazing
- Herbicide Options For Broad Leaved Weed Control In Seedling Chicory
- Agrochemical Free, Direct Sowing Culture Of A Paddy With Non Woven Fabric Mulch: Effects Of Sowing Time And Fertilizer Type
- Evaluation Of Within Crop Weed Management For Organic And Conventional Grain Production In South East Australia
- Ideas For Slug Control In Broadacre Crops
- Atrazine Management Guidelines For Central Queensland
- Profitable And Sustainable Perennial Grazing Systems: Searching For The Right Shrub Species
- Grazing Balansa Clover And Puccinellia Mixed Pastures On Saline Land
- Effect Of Harvesting Stage, Location And Genotype On Environmental Staining In Faba Bean (Vicia Faba L.)
- 4671_dhammuh
- Grazing Effects On The Retranslocation Of Assimilates During Grain Filling Of Wheat
- Food Safety Risks Posed By Pesticide Residues In Changing Cropping Systems
- Perennial Pastures For WA
- High Yielding Weevil Tolerant Maize Is Attainable For Subsistence Farmers In East Timor
- Vegetable Beetle, Gonocephalum Misellum (Blackburn), A Pest Of Canola In Southern Western Australia.
- Five Year Lucerne Pasture Improves The Growth Of The Following Canola Crop On A Sodosol With Dense Subsoil
- Overcoming Soil And Nutrient Constraints
- “Jota” Annual Sweet Clover (Melilotus Albus Medik.): A New Salt Tolerant Legume For The High Rainfall Zone Of Southern Australia.
- A New Generation Ripper To Reduce The Cost Of Removing Soil Compaction
- 4553_daviess
- Tolerance Of Wheat Genotypes To Subsoil Constraints In Southwest Queensland
- Seasonal Expression Of Subsoil Constraints In A Mediterranean Type Environment
- A New Standard For Electromagnetic Induction Mapping Of Soils For The Grains Industry
- Waterlogging Highlights Benefits Of Rhizobia In Faba Beans.
- Fate Of Nutrients And Heavy Metal Contaminants In Biosolids Applied To Contrasting Soils And Cropping Systems In Southeast Queensland
- Resistance Of Rapeseed (Brassica Napus L.) To Aluminium Apparent In Nutrient Solution But Not In Soil
- Addressing Poor Nodulation And Molybdenum Deficiency In Chickpea Through Seed Priming
- Simulating Historic Cropping Systems To Understand The Interaction Between Crop Choice And Deep Drainage.
- Dryland Salinity And Agronomy In South East Australia: Groundwater Processes Or Soil Degradation Due To Intensive Grazing And Cropping?
- Building Crop And Pasture Systems For Sustained Productivity And Profitability Systems And Prediction
- Perennial Wheat: A Paradox With Possibility
- Summer Dormant Temperate Grasses Are Productive And Persistent In The Medium Low Rainfall Cropping Region Of New South Wales
- EverGraze – Development Of Profitable And Sustainable Livestock Systems For The High Rainfall Zone Of Western Australia
- Nitrogen Balance Of Kale
- Developing Collaborative Planning Support Tools For Optimised Farming In Western Australia: A Methodology
- Rice Response To Organic Residues Management On Permanent Raised Bed System In The Semi Arid Tropics Of Eastern Indonesia
- APSFARM: A Whole Farm Systems Analysis Of Economic And Environmental Indicators Of Contrasting Farm Business Strategies In Central Queensland
- The Potential Of Winter Wheat Cultivars And Breeding Lines For Use In Dual Purpose (grain And Graze) Systems
- Are There Pre Sowing Indicators For Choosing In Which Seasons To Companion Crop Wheat Into Existing Lucerne ?
- Predicting Wheat Flowering Dates In Western Australia
- Growing More Grain: A First Cut At Estimating And Locating Potential Gains
- The Application Of Precision Agriculture Techniques To Assess The Effectiveness Of Raised Beds On Saline Land In WA.
- A Survey Of Perennial Pasture Establishment Practices In Southern NSW
- Effects Of Pasture Leys, Tillage System And Crop Frequency On C Dynamics, Soil Physical And Biological Properties
- Identifying Vegetable Lablab Types By Participatory Assessment: Panelists’ Perceptions Of Morphological Traits And Organoleptic Taste Assessment
- Do Seasonal Climate Risk Management Tools Address The Risk?
- Profitability And Cost Of Production
- Using Crop Modelling To Supplement Crop Trial Data For Land Capability Assessment In Cambodia
- Contribution Of Potential Yield, Drought Tolerance And Escape To Adaptation Of 15 Rice Varieties In Rainfed Lowlands In Cambodia.
- Flamenco A New Variety Of Sulla For Southern Australia
- Future Technologies In Agriculture
- Improving A Spatially Referenced Crop Simulation Model By Accounting For Salinity
- Spectral Response Of Nitrogen Fertilization In Cotton (Gossypium Species)
- Automatic Identification Of Soil Spectral End Members From Multi Temporal MODIS Imagery In The Western Australian Wheat Belt
- APSIM Barley Model – Adaptation Of A Wheat Model To Simulate Barley Growth And Development
- RAPSIM, A Statistical Toolbox For Analysing And Visualising Large Simulation Data Sets
- Seasonal Changes In Pasture Quality In Mediterranean Regions Of Australia
- The Australian National Agricultural Monitoring System – A National Climate Risk Management Application
- Managing Water In A Changing Environment
- The Role Of Groundwater Depth On The Hydrological Benefits Of Lucerne And The Subsequent Recharge Values
- Climate Change Impact On Field Crops And Adaptation Options In Southeastern Australia
- Measurement Of Dry End Soil Water Diffusivity On Intact Soil Cores
- Effect Of Water Deficit On Yield Of Pasja Forage Brassica
- Delivering Soil Water Information To Growers And Consultants
- 4572_hassanf
- The Effect Of Sorghum Row Spacing On Fallow Cover Distribution And Soil Water Accumulation In Central Queensland
- Investigation Into Partial Root Zone Drying In Cotton Cropping Systems
- Growth And Water Use Response Of Two Wheat Cultivars Exposed To Water Deficit At Four Growth Stages
- Limited Ability Of Native Pastures To Control Deep Drainage
- Can Variability In Subsoil Constraints Be Spatially Managed In The High Rainfall Zone Of SE Australia?
- Participatory Research: Adopting Agronomy
- Concurrent Papers
- 2008 14th AAC
- Concurrent Papers
- Achieving Change
- Perceptions Of Grain Growers Towards Annual Cropping In The High Rainfall Zone Of Southern Australia
- Evaluating Change In Knowledge And Practices To Combat Subsoil Constraints In North Eastern Australia
- Farmer Participatory Varietal Selection In Lao Lowland Rice Systems
- A Sociological Approach To The Participatory Development Of Agricultural Decision Support Systems
- Participatory Variety Selection Increases Adoption Of Modern Varieties By Subsistence Farmers In East Timor
- Agronomy Abroad
- Effect Of Farmyard Manure (FYM) And Inorganic Fertilizers On The Yield Of Maize In Wheat Maize System On Eroded Inceptisols In Northern Pakistan
- Soil Clay Contributing To Determination Of Growing Period In Rainfed Lowland Rice
- Forages For Agricultural Production And Catchment Protection In Eritrea
- Crop Diversification In Lowland Rice Cropping Systems In Cambodia: Effect Of Soil Type On Legume Production.
- Evaluating Use Of Remote Sensing For Identifying Management Strategies: Example For Small Plot Farmers During The Dry Season In Southern Bangladesh.
- A Preliminary Survey Of Cropping Systems In Tibet
- Developing A Strategy For Improved Crop And Animal Production In The Semi Arid Tropics Of West Timor
- Conservation Agriculture In A Subsistence Farming System: Lessons From On Farm Research In Gansu, China
- Herbicide Resistance Management In Phalaris Minor And Resource Conservation Technologies In Rice Wheat Cropping System In India – An Overview
- Expanding Production And Use Of Legume Inoculants In Myanmar And Vietnam
- Agronomy In The Landscape
- Paddock Scale Water Use In A ‘belt And Alley’ System With Oil Mallee Trees
- Effect Of Salinity On Seed Germination Of Wheat Cultivars
- Saltland Capability Assessment: Targeting Plants To Landscapes To Increase Profitability
- Farming Practices To Improve Water Quality In The Burdekin Region
- Precision Agriculture For Improved Environmental Outcomes
- Defining Tradeoffs Between Farm Profit And Natural Resource Indicators In Grain Grazing Systems
- Ecosystem Services In Agricultural Landscapes
- Agronomy As A Profession
- Assessing Yield Potential
- Relationships Between Growing Season Rainfall, Grain Yield And Yield Components Suggest That Wheat Requires One Ear/m2 Per Millimetre Of Rainfall To Achieve Its Water Limited Potential
- 5779_huntjr
- Estimating Grain Yield With The French And Schultz Approaches Vs Simulating Attainable Yield With APSIM On The Eyre Peninsula
- Modification To The French And Shultz Formula To Account For Soil Type And Within Season Rainfall.
- A National Benchmark For The Australian Wheat Industry: Accounting For Overlooked Climate Drivers
- Soil Nitrogen And Water Dynamics In Crops Following Perennial Pastures Under Drought Conditions.
- Biotechnology, Paddock Reality
- Do Sowing Rules Change For High Fruit Retention Transgenic Cotton?
- A Wheat Ideotype For The High Rainfall Zone Of South West Victoria
- Effect Of Early Moisture Deficit On Growth, Development And Yield In High Retention Bt Cotton
- Emerging Opportunities For Agriculture: Investigating Plant Adaptation By Characterizing Germplasm Collection Habitats
- 5858_chenu
- Development Of The Generic APSIM Cereal Template To Enhance Functionality Of Whole Plant Modelling
- Crop Agronomy
- Soil Borne Disease Suppression Of Rhizoctonia Root Rot In Soils Of The Eyre Peninsula, South Australia
- Wheat And Canola Water Requirements And The Effect Of Spring Irrigation On Crop Yields In The Central Murray Valley
- How Important Is Straw For Yield Of No Till Crops On Heavy Soils In The Low Rainfall Southern Mallee?
- The Effect Of Faba Bean Plant Population On Yield, Seed Quality And Plant Architecture Under Irrigation In Southern NSW.
- Farming System Impacts On Microbial Activity And Soil Organic Matter Dynamics In Southern Australian Mallee Soils
- Crop Growth
- Simulation Of Wheat Grain Yield Under Elevated CO2 Conditions In The Victorian Wimmera
- A Critical Analysis Of Osmotic And Ionic Effects Of Salinity In Two Barley Cultivars
- The Effect Of Elevated Carbon Dioxide On The Growth And Yield Of Wheat In The Australian Grains Free Air Carbon Dioxide Enrichment (AGFACE) Experiment
- Stem Nodes – The Central Building Blocks Of Growth And Yield In Field Pea.
- Crop Protection
- Site Specific Weed Management (SSWM) – Australian Potential With Annual Ryegrass (Lolium Rigidum L)
- Pre Emergent Herbicides For Managing Resistant Ryegrass Populations
- Wide Row Lupin (Lupinus Angustifolius L.) Cropping Systems May Sustain Lupin Production In Western Australia
- Improving Summer Weed Control In Dryland Cropping Systems Through Targeted Research And Extension
- Emerging Opportunities
- An Economic Analysis Of Perennial Cereal Crops In Australia
- Diffusive Gradients In Thin Films (DGT) As A Technique For Accurately Predicting Phosphorus Fertiliser Requirements
- Estimating Adaptive Capacity In Australian Farming Environments
- Using The Farm Scale System Model APSFarm To Improve Profitability Of Irrigation Cropping Enterprises.
- Assessing The Availability Of Crop Stubble As A Potential Biofuel Resource
- How Effective Are Fluid Phosphorus Fertilisers In The Wimmera And Mallee Region Of Victoria?
- Evaluating Systems
- Cotton Yield And Soil Carbon Under Continuous Cotton, Cotton Corn, Cotton Vetch Corn And Cotton Wheat Rotations
- Crop And Whole Farm Tools For Analysis Of Profit, Risk And Environmental Impact
- The Determinants Of South Australian Wheat Yield Increases
- Why Do We Persist With Static Equilibrium Whole Farm Economic Models?
- Farming In An Uncertain Climate
- A Dynamic Simulation Model For An Agricultural Catchment Driven By Markets, Farmers Attitude And Rainfall Change
- 5623_alexanderbm
- Shire Scale Impacts And Adaptation Options For Australian Cereal Crops Affected By Climate Change
- Examining The Value Of Dynamic Seasonal Forecasts In Managing Farm Level Production And Environmental Outcomes In A Variable Climate
- South Australian Farmers’ Concerns And Adaptation Options For Climate Change
- Paddock Change For Climate Change
- Managing Pastures
- Within Paddock Variation In Pasture Growth: Landscape And Soil Factors
- Pasture Characteristics Perceived By Farmers Of Western Australia In Relation To Adoption Of Annual Pasture Legumes
- Autumn Cleaning Ornithopus Compressus L. (yellow Serradella) Pastures: A Novel, Broad Spectrum Weed Control Option That Exploits Delayed Germination.
- Sheep Liveweight Production From Lucerne, Cocksfoot Or Ryegrass Based Pastures
- Perennial Pastures In The Wimmera Mallee
- Comparing The Water Use Efficiency Of Tropical Pasture Grasses And Legumes Used In Queensland’s Mixed Farming Systems
- Managing Site & Season
- Grower Evaluation Of Nitrogen Application Timing And Decision Tools In Uncertain Seasons.
- Comparison Of NDVI Seasonal Trajectories And Modelled Crop Growth Dynamics
- Farmer Case Studies On The Economics Of PA Technologies
- How Important Is Season Specific Or Soil Specific Wheat Agronomy In Southern Australia?
- The Role For EM Mapping In Precision Agriculture In The Mallee
- Does Spatial Variability Of Plant Available Water Capacity Cause Variability Of Grain Yield In The Victorian Mallee?
- Managing Soils
- The Long Term Consequences Of Soil PH Management On Crop Productivity And Profitability
- Sodium, Chloride, Clay And Conductivity: Consistent Relationships Help To Make EM Surveys Useful For Site Specific Management In The Mallee
- Are Reduced Acidification Rates A Feasible, Achievable Option For Future Agricultural Systems?
- Successful Integration Of Soil Nutrient Testing And Regional Acidity Monitoring Highlights The Opportunities For Soil Condition Assessment And Natural Resource Management
- Managing Subsoils
- Soil Compaction Is A Major Issue Operating Against The Development Of Sustainable Sugarcane Cropping Systems
- Simulating Seasonal Response Of Wheat Yield To Subsoil Amelioration In The Victorian Mallee And Wimmera
- 5838_mcbeathtm
- Subsoil Constraints Have Little Impact On Crop Growth In The Wimmera And Mallee During Dry Seasonal Conditions.
- Ripping Yarns: 25 Years Of Variable Responses To Ripping Clay Soils In South Eastern Australia
- Crop Production Responses To Soil Amelioration Treatments On An Infertile Sand Over Clay In South Australia
- New Grazing Options
- New Pastures And Forage Options
- Twin Sowing: A New Technique To Reduce The Costs Of Pasture Establishment.
- Current And Future Use Of Pasture Legumes In Central And Southern NSW – Results Of A Farmer And Advisor Survey
- Using Plastic Mulch To Increase Maize Silage Production In A Cool Climate With A Short Growing Season
- Dry Matter Yields And Nutritive Value Of Silage From Cereal And Pea Combinations
- EverGraze: The Productive Management Of Perennial Grazing Systems In Extreme Drought
- Benefits Of Small Seeded Annual Pasture Legumes In Pasture Mixtures For Variable Environments
- Pasture Production
- Herbage Production, Nitrogen Fixation And Water Use Efficiency Of Ten Annual Pasture Legumes Grown With And Without Lime On An Acid Soil
- Simulating Competition Between Cereal And Lucerne Grown In Mixture
- Developing Production Systems For The Newly Released Varieties Of The Biennial Forage Legume Sulla (Hedysarum Coronarium L.)
- Seasonal Variability Of Rhodes Grass Production In The Northern West Australia Wheatbelt.
- Lucerne Management Optimization – New Insights From Experimentation And Modelling
- Effectiveness Of Dairy First Pond Sludge As A Nutrient Source For Forage Crop Production
- Plant Nutrition
- Management Responses To Declining Potassium Fertility In Ferrosol Soils
- The Effects Of Maize Rotation On Soil Quality And Nutrient Availability In Cotton Based Cropping
- How Effective Are Fluid Phosphorus Fertilisers In The Wimmera And Mallee Region Of Victoria?
- SSP: Single Superphosphate, A Scenario Slowly Passing
- Rotations
- Crop Sequence To Manage Crown Rot In Cereals In South Eastern Australia
- Lucerne Production And Water Use In The Mallee
- Crop Sequences For High Productivity And Reduced Environmental Effects In New Zealand Dairy Forage Systems
- The Value Of Break Crops For Wheat
- Effect Of Lucerne Phase Duration In Cropping Systems In The High Rainfall Zone
- Achieving Change
- Conference Information
- Plenary Papers
- Poster Papers
- Agronomy In The Landscape
- Fallow Nitrate N Accumulation Rates In Central Qld Cropping Soils
- Plant Populations To Improve Yield Of Dryland Maize In Northwest NSW
- Impact Of Application Of Coal Combustion Products To Soil On Soil Characteristics, Concentration Of Elements In Plant Material And Crop Product Safety
- Dryland Cropping In Central Queensland Impacts On Soil Chemical Parameters.
- Effect Of Salinity On Seed Germination Of Wheat Cultivars
- Selection Of Wheat For High Early Vigour
- Row Spacing, Water Use, And Yield Of Wheat (Triticum Aestivum), Barley (Hordeum Vulgare) And Faba Bean (Vicia Faba)
- An Update On The Role And Agronomy Of Safflower In Southern Australia
- 5820_acunatl
- EM38 And Crop Soil Simulation Modelling Can Identify Differences In Potential Crop Performance On Typical Soil Zones In The Mallee.
- Simulation Of Field Pea Growth And Yield In Diverse Environments
- Resolving The Edaphic Conflict In Rice Wheat System
- Out Of Season Crops – What Are The Benefits For No Till Farming Systems?
- Primary Production Landscapes Of Victoria
- Organic Matter Fractions And Carbon Mineralisation In Perennial Pasture Leys On Erodible Silt Soil
- Biotechnology, Paddock Reality
- Increasing Grain Size And Reducing Screenings In Wheat Using A Tiller Inhibition Gene – Investigating Grain Morphology By Image Analysis
- Evaluation And Utilisation Of Pea And Faba Bean Germplasm From China.
- Variability In Structure And Function Of Sorghum Root Systems
- Plant Design Features That Improve Grain Yield Of Cereals Under Drought
- Can Large Kernel Size Increase Grain Yield In Sorghum?
- Genetic Diversity In Tropical Legumes: Cowpea (Vigna Unguiculata (L.) Walp.) And Lablab (Lablab Purpureus (L.) Sweet)
- The Role Of Genetic Solutions For Overcoming Subsoil Constraints To Lentil
- Emerging Opportunities
- Effectiveness Of Dairy First Pond Sludge As A Nutrient Source For Perennial Ryegrass Pasture Production
- Geo Referenced, Catchment Based Soil PH Monitoring In Three Agro Ecological Regions Of Western Australia.
- Australian Plant Genetic Resources Activities In 2006 And 2007
- Glycinebetaine And Salicylic Acid Application Improves The Plant Water Relations, Water Use Efficiency And Yield Of Sunflower Under Different Planting Methods
- Herbicide Resistance In Littleseed Canarygrass (Phalaris Minor) And Its Management
- Zero Tillage Wheat And Unpuddled Rice: The Energy, Labour And Cost Efficient Alternatives To Conventional Rice Wheat System
- Optimising The Quality And Yield Of Spelt Under Organic Production In SE Australia
- Farmer Focussed Research
- Effect Of Irrigation On Yield, Quality And Water Use Efficiency Of Indian Mustard (Brassica Juncea).
- Effect Of Nitrogen And Zinc Fertilisers On Yield And Protein Content Of Durum Wheat (Triticum Turgidum Var. Durum)
- Traditional Soil Fertility Management Strategies: Do They Conform To Recommendations In Organic Farming? A Case Study Of The Smallholder Farmers Of The Central Rift Valley Province Of Kenya
- Selection Of Widely Adapted Lowland Rice Varieties For Wet And Dry Seasons In Laos
- Effect Of Post Emergent Solid And Liquid Nitrogen On Grain Yield And Quality Of Bread And Durum Wheats
- Tillage And Nitrogen Management In A Wheat Maize Farming System
- Responses Of Bread And Durum Wheat To Agronomic Practices In Western Australia
- Evaluation Of A Proximal Vision Data Acquisition System For Measuring Spatial Variability In Lettuce Growth
- Development Of New Lentil Varieties In Bangladesh
- Survey Of Soil Borne Disease Suppression To Rhizoctonia Solani In Low Rainfall Farming Systems On Upper Eyre Peninsula, South Australia.
- Changed Farming Practice In Central Queensland Grain Farming Systems.
- An Evaluation Of Two New Vetch Cultivars In The Wimmera Region Of Victoria
- How Much Fuel Does Your Farm Use For Different Management Operations?
- Variable Response To Phosphorous Fertilisers In Wheat And Chickpea Crops In Central Queensland
- Responses Of Biomass, Chlorophyll And Macro And Micronutrient Uptake Of Rice (Oryza Sativa L.) To Organic And Chemical Fertilisers
- The Effects Of Zn And K Fertilisers On Agronomical Characteristics And Zn, Fe And P Accumulation In Two Wheat Cultivars In Calcareous Soils
- Grain Yield Of Heritage Wheat Varieties In South Australian Mallee, 2001.
- A Standards Based Approach To Farmers And Scientists Sharing Information Via Electronic Networks
- Using Local Case Studies In The High Rainfall Zone Of South Eastern Australia To Generate Grower Confidence In Computer Model Outputs
- Thirty Years Of Change In South Australian Broadacre Agriculture
- Optimising Spray Efficacy In Summer: Delta T Red Alert System
- Long Term Performance Of Intensive Cereal Based Cropping In The Mallee
- An Index For Evaluating Crop Production Variability From Remote And Proximal Sensor Data
- The Influence Of Agronomic Management On Seed Size Attributes In Kabuli Chickpea.
- Effects Of Plant Density On The Yield Of Field Pea And Faba Bean Varieties Across Southern And Central NSW—preliminary Findings
- Participatory Variety Selection Increases Adoption Of Modern Varieties By Subsistence Farmers In East Timor
- Farmer’s Participatory Research On Integrated Farming System
- Managing Water Soluble Carbohydrates And Fibre In Oaten Hay
- Farming In An Uncertain Climate
- Possible Impact Of Climate Change On Rice Production In The Gangetic West Bengal, India
- Screening Rice Genetic Resources For Cold Tolerance At Different Growth Stages
- Model Based Analysis Of Maize Plastic Response To Water Stress For Screening Traits In Improving Breeding Efficiency
- Revising Crop And Architectural Models Of Maize Canopy Development For Rainfed Cropping Conditions
- Using Modelling And Agronomy To Improve Reliability Of Maize As A Rainfed Crop In Northern New South Wales And South Western Queensland
- Modelling The Effect Of Temperature, Soil Moisture And Photoperiod On Growth And Development Of Bambara Groundnut (Vigna Subterranea (L.) Verdc.): BAMGRO Model
- Interactions Between Wheat Varieties And Fungicide Applications In South West Victoria
- Interpolating Leaf Area Index Between Destructive Sampling Dates Using Remote Sensing In The Australian Grains Free Air Carbon Dioxide Enrichment (AGFACE) Experiment
- Nitrous Oxide Emissions From Dry Land Wheat In South Eastern Australia
- The Development Of A Model In APSIM For The Simulation Of Grazing Oats And Oaten Hay
- Climate Change – Identifying The Impacts On Soil Health In Victoria.
- The Australian Grains Free Air Carbon Dioxide Enrichment (AGFACE) Experiment – Specifications And Scope
- Talking About The Weather: APSIM, Climate Change And Grain Farmers On The Upper Eyre Peninsula, SA.
- Managing In The Face Of Uncertainty: Is There Anything Farmers Can Learn From Finance?
- Impact Of Soil Properties And Climate Change On Nutrient Use Efficiency In Farmer’s Fields
- Establishment And Root Morphology Of Eight Diverse Lucerne Populations In A Low Rainfall, South Australian, Environment
- Dry Matter Production And Partitioning In Rice In Response To Elevated Temperature
- Climate Change Risk Assessment Associated With Wheat Production Systems In Southern NSW
- Integrated Systems
- The Use Of Ley Pastures To Arrest Soil Fertility Decline In Central Queensland.
- Sustainable Sheep Production In Mallee Farming Systems
- Dual Purpose Canola – Recent Results From Southern NSW
- Development Of An Australian Pasture Selection Tool Using Lucid™
- 5705_nidumoluub
- Trade Offs For Ratooning Sorghum After Harvest To Provide Forage For Grazing
- When Is It Profitable To Sacrificially Graze A Wheat Crop? A Simulation Analysis Across Australia’s Cropping Belt
- Effect Of Grazing On Cereal Forage DM Yields For Whole Crop Silage
- Leaf Appearance Rate And Tiller Development Rate In Pre Grazing Growth Of Wheat And Oats Under Dual Purpose (grain And Graze) Management
- Assessing The Suitability Of Several Tropical Pasture Species For Use As Leys Within Cropping Systems In Southern Queensland
- Harvestability And Potential Seed Production Of Cullen Australasicum
- Barley Benefits From Grazing
- 6210_deangj
- The Integration Of Novel Forage And Grain Systems In Southern Australia
- Agronomy In The Landscape
- Concurrent Papers
- 2010 15th AAC
- Climate Change
- Adaptation
- Fast Tracking The Adaptation Of Grain Production Systems To A Changing Climate Using A Participatory Action Research, Development And Extension (PARD&E) Process
- Using Degradable Polymer Film (DPF) To Mitigate The Impacts Of Climate Variability On Agricultural Production In Low Rainfall Areas
- Role Of Seed Priming In Improving Wheat Performance In A Changing Climate
- Using A Participatory Action Research, Development And Extension (PARD&E) Process To Adapt Grain Production Systems To A Changing Climate
- Adapting To Climate Change In Australian Farming Systems: A Network For Applied R&D.
- CO2
- The Size Of Free Air Carbon Dioxide Enrichment (FACE) Rings Affects Overall System Performance: An Australian Experience
- Future Effects Of Elevated CO2 On Wheat Production – An Overview Of FACE Research In Victoria, Australia
- Modelling The Growth And Grain Yield Response Of Wheat Crops Under Free Air Carbon Dioxide Enrichment At Horsham And Throughout Victoria, Australia
- N Dynamics Under Elevated Carbon Dioxide In The Australian FACE Experiment
- Intraspecific Variation Of Growth And Yield Response Of Wheat To Elevated CO2 In Australian Grains Free Air Carbon Dioxide Enrichment (AGFACE)
- Future Farming
- Mapping Potential Biofuel Crops In The Agricultural Landscape Of Victoria, Australia
- The Search For Alternative Perennial Pasture Legumes Adapted To The Changing Climatic Conditions Across Tasmania’s Low To Medium Rainfall Region
- A Survey Of The Current And Future Use Of Lucerne, Chicory And Perennial Grasses In Pasture Crop Rotations In Southern New South Wales
- Insect Pathogen Crop Dynamics And Their Importance To Plant Biosecurity Under Future Climates: Barley Yellow Dwarf Virus And Wheat – A Case Study
- Securing Our Food Supply Over The Next 10,000 Years
- High Temperature
- Future Elevated Carbon Dioxide Levels Largely Overcome The Impact Of Hotter And Drier Conditions On Wheat Growth In The Victorian Wimmera
- Characterising The Risk Of Heat Stress On Wheat In South Australia: Meteorology, Climatology And The Design Of A Field Heating Chamber
- Impact Of High Temperature On The Reproductive Stage Of Chickpea
- Effects Of Elevated CO2 And Irrigation On Gas Exchange And Water Relations Among Two Wheat Cultivars
- Prediction
- Generating Daily Future Climate Scenarios For Crop Simulation
- Using Temporal And Spatial Analogues To Consider Impacts And Adaptation To Climate Change In The South Australian Grain Belt
- Impact Of Changing Climate On Cereal Productivity In Queensland
- Impact Of Climate Change On Wheat Yields In Western Australia. Will Wheat Production Be More Risky In The Future?
- Is There A Value In Having A Frost Forecast For Wheat In The South West Of Western Australia ?
- Impact Of Climate Change On Tasmanian Dryland Pasture Production
- Seasonal Rainfall Forecasts As An Adaptation Strategy For Climate Change
- Adaptation
- Crop Production
- Dual Purpose Crops
- Development & Herbicide Management
- Relative Tolerance Of Microlaena Stipoides To Glyphosate Herbicide
- Cereal Cultivars Show Differential Tolerance To In Crop Herbicides In South Australia
- The Strategic Use Of Chlorsulfuron In The Seed Production Of Medicago Littoralis Cv. AngelA
- Cold Tolerance Screening For Cotton Cultivars Using Germination Chill Protocols
- Cold Tolerance Of Rice Cultivars At The Booting Stage Can Be Improved By High Water Temperature During Vegetative Growth
- Field Evaluation Of Sensitivity Of Wheat To High Temperature Stress Near Flowering And Early Grain Set
- Relationship Between Seedling Ages, Total Leaf Number And Leaf Appearance Rate Of Transplanted Rice
- Is There A Value In Having A Frost Forecast For Wheat In The South West Of Western Australia ?
- Evaluation Of The Frost Tolerance Of Triticale Varieties And Other Winter Cereals At Flowering
- High Rainfall Zone
- Using Raised Beds To Reduce Waterlogging Of Pastures Pasture Productivity Effects
- Using Raised Beds To Reduce Waterlogging Of Pastures Impact On Runoff And Soils
- Adoption Of Practices And Technologies On Australian Grain Farms
- Yield Formation Of Wheat In The High Rainfall Zone Of South Western Australia
- Effect Of Time Of Sowing On Phenology Of Cereals Grown In A Temperate Environment In South Eastern Australia
- Providing High Rainfall Zone Growers With Confidence In New And Alternative Management Practices
- Phase Durations In Wheat Will Need To Be Optimised For Crops In The High Rainfall Zone Of South West Victoria To Achieve Yield Potential
- Improving Crop Water Use Efficiency In High Rainfall Zone (HRZ) Victoria, Through Appropriate Practice Change To Overcome Subsoil Limitations
- Pulse Crops Improve Productivity Of Crops In Rotations Involving Various Residue Management Strategies In The High Rainfall Zone Of Western Victoria – A Case Study Example For Grower Decision Support.
- Simulating Leaf Area Duration To Predict Yield Response To Foliar Fungicide In Wheat And Barley
- Spatial And Temporal Variability In Organic Carbon Observed In Soil Under Lucerne Pastures
- Intercrops, Cover Crops, Companion Crops
- Screening Native C4 Pasture Genotypes For Shade Tolerance In Southern Brazil
- Estimated Long Term Trends In Agronomic Strategies For Mitigating Competition In Companion Cropping Systems
- A Comparison Of Lucerne And Chicory Based Intercrops And Competition Effects
- A Comparison Of Land Equivalent Ratios And Water Use In Lucerne And Chicory Based Intercrops
- Development Of Sustainable Legume Production In Rice Based Farming Systems In Cambodia
- Irrigated Crops
- Can Planting Date And Cultivar Selection Improve Resource Use Efficiency Of Australian Cotton Systems?
- Evaluation Of Irrigation And Nitrogen Management Strategies For Wheat Production On The Darling Downs
- Evaluation Of The Surface Renewal Method To Estimate Crop Evapotranspiration Of A Cotton Field
- Cultivation Methods Of Paddy: New Opportunities For Bangladesh
- Opportunities To Reduce The Impact Of Water Logging On Cotton
- Legume Agronomy
- A New Highly Virulent Bluegreen Aphid Causes Severe Damage In Previously Tolerant Pasture And Grain Legumes
- Genotype à Environment à Herbicide Interaction In Narrow Leafed Lupin (Lupinus Angustifolius L.)
- Improving The Reliability Of Early Sown Pulses In South Eastern Australia
- Measuring The Impact Of Bacterial Blight (Pseudomonas Syringae Pv. Syringae) On Production Of Australian Field Pea Varieties.
- The Importance Of Inoculant Type Or Application Rate For Nitrogen Fixation By Grain Legumes In Cropping Systems Of South Eastern Australia
- Nitrogen And Phoshorus
- Sulphur Enhanced Triple Superphosphate As A Substitute For Single Superphosphate
- Phosphorus Use Efficiency In Wheat And Barley
- Expanding The Use Of Diffusive Gradients In Thin Films (DGT) For Assessing Phosphorus Requirements Of Different Crop Types
- Effect Of In Season Moisture On Carbon, Nitrogen And Phosphorus In Alkaline, Semi Arid Soils
- The Farm Gate Phosphorus Balance Of Sheep Grazing Systems Maintained At Three Contrasting Soil Fertility Levels
- Nutrient Management
- New Tools For Real Time N Sensing Of Cereal Crops
- Developing A Searchable National Database Of Grain Soil Test Crop Response Information For Australia
- Combating Food Shortages Through Fertilizer Management Under Improved Planting Practice
- Two SALTs Simulation Assisted Learning Tools For N Fertiliser Rate Calculation In The Northern Grains Region
- ‘NBudget’ – A Nitrogen Management Tool For Cropping Systems
- Improved Modelling Of Manure Mineralisation Through New Methods For Characterising The Carbohydrate Pools
- Improved Prediction Of Wheat Yield Response To Nitrogen Curves
- Physiology And Breeding
- Evaluation Of Perennial Wheat Germplasm In An Australian Environment
- The Contribution Of Remobilization Of Storage Materials In Wheat Yield As Affected By Potassium Iodide
- Increased Stability Of Kernel Weight Under Drought Through Selection Of A Reduced Tillering Gene In Wheat
- High Insect Protection Of GM Bt Cotton Changes Crop Morphology And Response To Water Compared To Non Bt Cotton.
- Traits For Perennial Wheat Adaptation In Australia
- Plant Density
- Determining Physiological Cutout In Ultra Narrow Row Cotton
- The Impact Of Row Spacing On The Growth And Yield Of Lentil Cultivars In Southern Australia.
- Yield Response Of Kabuli And Desi Chickpea (Cicer Arietinum L.) Genotypes To Row Spacing In Southern Australia
- Variation In The Response Of Canola Cultivars To Changes In Row Spacing
- Agro Physiological Traits And Dry Matter Accumulation Of Corn Grown Under Varying Row And Plant Spacing
- Patterns Of Dry Matter Accumulation And Grain Yield Of Three Corn Hybrids Under Varying Planting Density/row Spacings
- Precision Agriculture
- Farmer Perspectives Of Precision Agriculture In Western Australia: Issues And The Way Forward
- Assessing The Whole Farm Benefits Of Controlled Traffic Technology
- Matching Inputs To Soil Potential In Low Rainfall Cereal Systems Of The Upper Eyre Peninsula
- Precision Agriculture For Pasture, Rangeland And Livestock Systems
- Crop And Pasture Sequence Decisions
- The Effect Of Brassica Species On Rhizoctonia Solani Disease Of Barley In Low Rainfall Farming Systems On Upper Eyre Peninsula.
- Making Better Decisions About Crop Rotations In Low Rainfall Environments: Should Stored Moisture And The Timing Of The Seeding Opportunity Influence This Decision?
- The Effect Of Soil Amendments And Soil Structure On Minimizing Constraints Of Lowland Soils On Growth Of Mungbean And Peanut Under Glasshouse Condition
- A Simple, Self Adjusting Rule For Identifying Seasonal Breaks For Crop Models
- Modelling The Profit And Risk Outcomes Of A Range Of Soil Water Thresholds And Cropping Area Intensities – A Whole Farm Approach
- Soil Biology And Nutrition
- Soil Water And Water Use Efficiency
- Interactive Effects Of Root Diseases And Drought On Water Use Efficiency Of Wheat
- Production, Water Use And Water Use Efficiency Of Short Term Ley Legumes In Southern Queensland
- Improving Winter Crop Productivity Through Increased Capture And Storage Of Summer Fallow Rain
- Benchmarking Farm Water Use Efficiency In Eastern Australian Dryland Cropping Systems
- The Effect Of Initial Soil Water On Grain Crop Yield And Yield Variability In North Eastern Australian Dryland Cropping Systems
- Benchmarking Wheat Water Use Efficiency In Tasmania
- The Effects Of Root Angle On Root Growth And Yield Of Wheat In The Australian Cereal Belt
- A Rapid Method For Estimating The Plant Available Water Capacity Of Vertosols
- Benefits Of Increased Soil Exploration By Wheat Roots
- Variable Soil Water Accumulation Under Fallow Management: Explanation Using A Pulse Paradigm
- How Much Is Enough: The Steep Final Steps To Extensive No Tillage Cropping In Australia
- What Influence Does Summer Rainfall And Stubble Retention Have On Wheat Production In Southern Farming Systems?
- Subsoil Constraints
- Impacts Of Simulated Subsurface Soil Compaction On Soil Properties And Barley Growth In Canterbury, New Zealand
- Phalaris And Lime – Improving Productivity On An Acidic Soil In A Drought Prone ‘high Rainfall’ Environment
- Removal Of A Subsoil Constraint When Does It Pay?
- On Farm Assessment Of Sub Soil Salinity And Sodicity Constraints To Barley Production In Southern Australia
- Integrated Weed Management
- Evaluation Of Integrated Weed Management Package For Soybean
- The Economic Benefits Of Glyphosate Tolerant Canola And Lupin In Controlling Annual Ryegrass (Lolium Rigidum)
- Stability Of Competitive Ability In Wheat Genotypes Across Different Weed Species
- Barley Grass, An Emerging Weed Threat In Southern Australian Cropping Systems
- Density Dependent Interference Of Emex Australis In Wheat At Different Sowing Times
- Farming Systems
- Animal Production And Feed Quality
- Energy Balance And Resource Use
- International Crop Pasture Systems
- The Use Of Forage Legumes In Cereal Cropping Systems Of Eastern Indonesia
- Improving Grain Yield And Quality Of Wheat Grown In Haryana (NW India)
- East India Plateau – Basket Case, Or Future Food Basket?
- Energy Balance Of Integrated Crop Rangeland Livestock Production Systems In Eastern Gansu, China
- Integrating Wheat Into The Rice Based Farming Systems Of Southern Bangladesh
- Agronomic Performance Of Landrace And Certified Seeds Of Maize In West Timor, Indonesia
- Nutrient Loss And Water Quality
- Participatory Research, Extension And Teaching
- Proofing Participatory Research
- The Development And Delivery Of A New Diploma Of Agronomy Course
- On Farm Participatory Research Investigating The Use Of Covercropping For The Establishment Of Perennial Pastures In Mixed Farming Enterprises
- Two Levels Of Information Are Required To Assist Extension Of Liming In Western Australia
- Simulation And Decision Support
- Integration Of A Pasture Model Into APSIM
- The Evolving Role Of Crop Modelling In Agronomy Research
- Is There A Better Way To Present Simulation Data And Probabilities?
- Developing Grass Curing Algorithms For Decision Support Tools
- Simulating Perennial Crops With The APSIM Plant Module – A Study With Lucerne
- Flowering Calculator: Parameters For Predicting Flowering Dates Of New Wheat Varieties In Western Australia
- Using Industry Information To Obtain Insight Into The Use Of Crop Rotations In The Western Australian Wheat Belt And Quantifying Their Effect On Wheat Yields
- Validating The GRAZPLAN Pasture Model For Native Grasslands Of The Monaro Region
- Bio Economic Modelling Of Beef Grazing Enterprises To Inform Sustainable Grazing Management Of Northern Australian Grazing Lands
- Soil Quality Carbon And Compaction
- Combined Effect Of Soil Moisture And External Loads On Soil Compaction
- Balancing Soil Carbon Quality And Quantity For Sustainable Agriculture
- Explaining Variability In Soil Carbon Stocks Based On Farm Management Factors
- Impact Of Landuse On Profile Distribution Of Fine Root Biomass In NSW, Australia
- Will Stubble Management And Stubble Load Affect Soil Organic Carbon Under Cropping In The High Rainfall Zone Of Victoria?
- Soil Science On The Ground
- Model Simulations Of Carbon Storage And Fluxes In Native Perennial And Temperate Grass Based Pastures
- Pastures And Forage
- Dryland Perennials
- The Effect Of Lime Application To An Acid Soil On Perennial Grass Establishment
- Climatic Conditions Associated With Successful Perennial Grass Recruitment Events In South Eastern Australia
- Sulla (Hedysarum Coronarium) Production Sown With Cover Crops
- Phalaris And Cocksfoot Prove Superior To Tall Fescue In Two Drought Prone Environments Of Southern NSW
- Fork In The Road: Forage Shrub Plantings By Crop Livestock Farmers In A Low Rainfall Region Of Southern Australia
- Forage Crops Brassicas
- Forage Crop Production
- Production And Nutritive Value Of Alternative Annual Forage Crop Options In A Rainfed Region Of Western China
- Determinants Of Productivity In Cropping Sequences For Dairy Systems In Canterbury, New Zealand
- Predicting Yield And Biomass Nitrogen Of Forage Crop Rotations In New Zealand Using The APSIM Model
- Using Growth And Dry Matter Estimates To Devise Year Round Forage Systems For The North West Slopes Of New South Wales
- Dry Matter Yield And Nutritive Value Of Forage Crops Under Different Rotations On The Longdong Loess Plateau, China
- Legumes And Broadleaf
- Chicory Increases The Level Of N Fixation By Companion Legumes
- Tedera Out Yields Lucerne And Perennial Ryegrass Five Months After Sowing
- Improving The Nodulation Of Regenerating Stands Of Messina (Melilotus Siculus) In Saline Soils
- Growth And Development Of Lotus And Trifolium Species Under Saline And Waterlogging Conditions
- Estimated Symbiotic N2 Fixation By Annual Legume And Lucerne Pastures On 2 Vertosols With And Without Applied Gypsum.
- Lucerne Systems
- Impact Of Including Lucerne And Other Perennials In Crop Livestock Enterprises In Southern NSW
- The Potential Of Summer Growing Grasses To Persist And Produce Out Of Season Forage In The Victorian Mallee.
- Farmers’ Experience With Lucerne In Western Australia
- Opportunities And Challenges To Increased Lucerne Adoption In New South Wales, Australia
- Physiology And Breeding
- A Description Of The Plant Structures Of Microlaena Stipoides, A Grass Species Forming Stolons And Rhizomes
- Effect Of Planting Date And Daylength On Establishment Of Dorycnium Hirustum
- Comparison Of Functional Food Components In Grain Of High Anthocyanin GM And Non GM Rice
- Screening For Drought Resistance Among A Large Number Of Australian Green Couch Grass (Cynodon Spp.) Ecotypes During Canopy Establishment
- An Early Maturing Trifolium Subterraneum L. Var. Yanninicum With Improved Agronomic Traits
- Assessment Of Root Growth By Perennial Pasture Grasses In An Acid Soil Using Novel DNA Based Methods
- Spatial Management
- Pasture Response To Superphosphate Application In Topographically Diverse Landscapes Of The Central Tablelands And Monaro Regions Of New South Wales
- Factors Affecting Pasture Growth In A Topographically Diverse Native Perennial Grass Pasture On The Central Tablelands Of New South Wales
- Using High Resolution Landscape And Soils Data To Understand Spatiotemporal Variability In Net Pasture Productivity As Derived From Low Spatial Resolution Remote Sensing.
- Spatio Temporal Movement Of Livestock In Relation To Decreasing Pasture Biomass
- Satellite Remote Sensing As A Tool To Quantify The Area Of Lucerne At A Paddock Scale In The Fitzgerald River Catchment Of Western Australia
- Uneven Nutrient Distributions In Hillside Paddocks Indicate Potential Need For Variable Rate Fertiliser Applications To Pastures
- Integrated Weed/Insect Management
- Dryland Perennials
- Plenary Papers
- Climate Change
- 2012 16th AAC
- Agriculture
- Crop Yields And Food Security: Will Yield Increases Continue To Feed The World?
- Economics, Productivity And Natural Resources In Agricultural Systems
- The Value Of A Boundary Organization In Mediating Knowledge On Sustainable Farming Systems
- Sustainable Grains Production Course – Building Grains Industry Futures.
- Australia’s Declining Crop Yield Trends I: Donald Revisited.
- Serious Games To Explore Uncertainty Of Future Farms
- Genetically Engineered Forage Crops: New Zealand Public Attitudes
- Managing The Key Risks Of Farming; Climate And Commodity Price Variability
- Sustainable Crop Intensification: Capturing Opportunities In A Highly Variable Climate With Relay Crops
- 2012 Authors
- Breeding
- Investigation Of Gene Expression Underpinning Partitioning Of Seed Storage Compounds In Legumes
- Yield Components Of High Yielding Australian Cotton Cultivars
- Genotype By Environment (GxE) Interactions For Root Depth Of Wheat: Associations And Implications
- Sugarcane For Water Limited Environments: 3.Transpiration Efficiency Of Commercial And Wild Relatives Of Sugarcane
- A Model For Predicting Milling Yield In Wheat
- Rice Germplasm Selection And Production Systems For The Northern Territory.
- Grain Protein Concentration Of Several Commercial Wheat Varieties
- High Temperature Effects On Development And Floret Sterility Of Diverse Sorghum Lines
- Persistence Of Pasture Legumes In Southern And Central Queensland
- Potential Use Of New Generation Annual Pasture Legumes In Crop Pasture Rotations In Central And Southern NSW
- Characterisation Of Drought Stress Dynamics In European Maize Crops
- Field Assessment Of Pre Harvest Sprouting Of Wheat Varieties In Western Australia
- Climate Change
- The Challenge Of Engaging With Farmers About The Impacts Of, And Their Adaptation To, Climate Change
- Know The Factors That Affect The Usage Of CO2 In A FACE System And Adjust Accordingly
- Predicted Frequency Of Wet And Dry Soil Conditions In Tasmanian Dairy Regions Under Future Climate Scenarios
- Soil And Pasture Responses To Poultry Litter Biochar Combined With Nitrogen Fertiliser On A Degraded Red Vertosol In Tamworth, NSW Australia
- The Potential For Inhibitor Coated Nitrogen Fertilisers To Reduce Agricultural Fertiliser N Losses
- Use Of Nitrification Inhibitors To Reduce N2O Emissions From Southern Pastures
- Effect Of Elevated CO2 And N Level On Growth Of Wheat And Field Pea
- Potential Biodiesel Crops For Marginal Land In Australia
- Adaptation Strategies To Climate Change For New Zealand Cropping Rotations
- Genetic Resources And The Challenge Of Climate Change
- Paired Sites To Engage Farmers And Understand Soil Organic Matter And Carbon
- Particulate And Mineral Associated Organic Carbon Fractions As Influenced By Corn Residue Incorporation And Simulated Tillage
- Timing Of Autumn Breaks And Length Of Springs In Tasmanian Dairy Regions Under Future Climate Scenarios
- Heat Waves And Wheat Growth Under A Future Climate
- Effects Of Elevated CO2 On Vitamin E Concentrations In Grains Of Two Wheat Cultivars – A FACE Study
- Participatory Climate Risk Assessment With Dryland Farmers
- Complex Feedbacks Can Make Agronomic Management Of Greenhouse Gas Emissions Difficult In Subtropical Agricultural Systems.
- Mapping The First Summer Rains In The Northern Agricultural Region Of Western Australia
- Mineralisation Of Chicory And Lucerne Residues Under Field Conditions
- Opportunities To Sequester Carbon In Soil: Management Of Perennial Pastures
- Tracking Climate Adaptation Of Growers In The Victorian Wimmera Through Varying Seasonal Conditions
- Historical Trends In Rainfall And Temperature In Queensland’s Mixed Farming Zone
- Scenarios Of Future Climates At The Sub Regional Scale In Queensland’s Mixed Farming Zone
- Impact Of Climate Change On Wheat Production And Profitability In Queensland
- Can Agronomic Management Mitigate N2O Loss From High Rainfall Cropping Systems?
- Greenhouse Gas Emissions From Legume And Grass Pastures Prior To Cropping.
- Climate Change: What Mixed Farmers In Queensland Really Think And Their Strategies To Manage It.
- Biochar Can Enhance Soil Fertility And Reduce Greenhouse Gas Emissions
- Drought Experienced By Australian Wheat: Current And Future Trends
- Weather Certificates And Their Use To Mitigate Adverse Weather Conditions
- The Impact Of Climate And Technology On Australian Grain Yields.
- Crop Development
- Allometric Relationships Of Canopy Organ Development In Rice And Maize
- Maize Canopy Development In Response To Increasing Plant Population Density
- Vegetative Development Of Four Annual Clovers
- Regrowth And Grain Recovery Of Spring Wheats Following Early Defoliation
- Canopy Development And Radiation Use Efficiency Of Four Forage Brassica Crops
- 8068_riffkinpa
- 8120_peakeas
- Effects Of High Temperature At Different Developmental Stages On The Yield Of Chickpea
- Mechanisms Of Drought Response In Summer Forage Brassicas
- Optimising Flowering Time, Phase Duration, HI And Yield Of Milling Wheat In Different Rainfall Zones Of Southern Australia
- Effects Of Plant Growth Regulators That Reduce Stem Height On Yield Of Wheat In Southern Australia
- The Effect Of Time Of Sowing On Phenology And Yield Of Chickpeas At Trangie Central West, NSW, 2011
- Root Development Of Rice Under Flooded And Aerobic Conditions
- The Influence Of Development On Early Growth In Wheat
- Crop Production
- Rice In The Ord: Opportunities And Threats
- Wheat Time Of Sowing And Cultivar Selection Can Be Used To Manage Risk In Low Rainfall Southern Australia.
- Break Crops Can Be Grown In The Mallee With A Low Risk Of Wind Erosion
- Grain Size Distribution: Computation, Interpretation And Utilisation For Minimising Small Grain Screenings In Cereals
- Improving Reliability Of Maize Production In Variable Rainfall Environments Of The North East Region.
- Break Crops Pay In The Victorian Mallee
- The Potential For Thin Biodegradable Film In The Australian Cotton Industry
- Soil Compaction Under Cotton Pickers: Preliminary Results
- Improved Crop Production In Response To Intensively Grazed Improved Legume Pastures
- An Alternative Storage Technique For Maintaining Maize Seed Quality In West Timor
- Frost Risk Associated With Growing Maize For Silage On Tasmanian Dairy Farms
- Break Crops Can Improve Wheat Production In The Mallee
- Diversification From Cropping Into Mixed Crop Livestock Systems – The Sustainability Risks Posed By Hay Removal From Pasture Or Forage Blocks
- Seeding Rate Responses Of New Durum Cultivars In Southern Australia.
- Expanded Row Configuration Options For Australian Rain Fed Cotton
- Assessment Of The Degree Of Impact Of Factors Affecting Micronaire In Cotton
- Using A Crop Model To Characterise The Developmental Phenotype Of Different Wheat Varieties.
- Integrated Agronomic And Economic Analysis Of Fodder Options For Tibetan Farming Systems
- Grazing Spring Variety Cereal Crops Reduces Supplementary Feeding In Mixed Cropping And Sheep Farms
- Getting N Management Right To Meet Malting Specifications For Barley
- Lodging Management For Commander Barley
- Decision Support Systems (DSS) –Where Success Is Failure Of Continued Use
- Scoping Early Sown Canola In Western Australia
- Canola Growth And Development In Central Western NSW
- The Influence Of Row Spacing On Grain Yield In Drill Sown Rice
- Rice Grain Yield A Comparison Between Direct Seeding And Transplanting In Lao PDR
- The Effect Of Sowing Depth On The Establishment Of Several Commercial Wheat Varieties
- The Influence Of Planting Date, Sowing Depth And Soil Type On Chickpea Production With No Tillage In Northern New South Wales
- Sequence Effects On Cropping System Productivity And Profitability In Two Environments In Western Australia
- VarietyChooser: The Flexible Cropping Management Tool
- Soil And Crop Management To Increase Income Of Rice Farmers In Aceh, Indonesia
- The Profitability Of Dryland Forage Crops: A Modelling Analysis In Central Queensland.
- Effects Of Row Spacing And Row Placement On Grain Yield In A Sorghum/wheat Sequence Under High Rainfall
- Achievable Sorghum Yields Under Favourable Conditions In North Western NSW.
- The Gap Between Yield Potential And Reality For Continuous Wheat In Wet Years In A Dry Environment.
- Potential For Dual Purpose Crops In Australia’s Northern Crop Livestock Regions
- Disease
- Grazing Barley Controls Early Foliar Diseases, Has Manageable Impacts On Malting Barley Grain Quality But Suffers A Yield Penalty.
- The Impact Of Crop Rotation And Nutrition On Rhizoctonia Disease Incidence In Cereals On Grey Calcareous Soils Of Upper Eyre Peninsula
- A Wet Summer Followed By A Wet Winter, A Recipe For Severe Take All Damage To Cereal Crops In Cereal Dominant Rotations
- Enhancing Plant Disease Prediction Using Leaf Wetness Forecasting
- Rhizoctonia Root Rot Suppression In An Alkaline Calcareous Soil From A Low Rainfall Farming System
- Seed Impurity Undermines Stripe Rust Resistance
- Nutrition
- Effect Of Rice Husk Biochar And Nitrification Inhibitor Treated Urea On N And Other Macronutrient Uptake By Maize.
- The Impact Of Lime And Gypsum On Pasture Composition In A Grass/legume Mixture Grown On A Shallow Acidic Loam
- Nutrient Use Efficiency In Current And Redundant Cotton Cultivars
- Early Growth Of Field Peas Under Saline And Boron Toxic Soils
- Seasonal Mobility Of Boron And Salt In Relation To Rainfall Across The Growing Season
- Break Crops For Disease And Nutrient Management In Intensive Cereal Cropping
- Wheat Grain Nutrient Concentrations For South Eastern Australia
- Soil Test Values And Nutrient Balances From A Long Term Fertilizer Experiment.
- Spatial And Temporal Trends Of CSBP Soil And Plant Analysis Data In Shires Of Western Australia
- Increasing Complexity In Nutrient Management On Clay Soils In The Northern Grain Belt – Nutrient Stratification And Multiple Nutrient Limitations
- Sodium Chloride Induced Salinity Reduces The Microbial Processes In Soil
- Sub Soil Constraints Affect Responsiveness Of Wheat To Applied Phosphorus
- Why Crops Fail To Produce Grain Yield Responses To Phosphorus Fertilisers?
- Can Soil Specific Inputs Of N Fertiliser Be A Risk Reducing Strategy?
- Lucerne (Medicago Sativa L.) Dry Matter, Root Growth And Nodulation In Response To Lime And Phosphorus In An Acid, High Country Soil In New Zealand
- Knowledge Is Power In Soil Acidity Management
- Wheat Yield Benefits From Fallow Due To Stored Soil Water And Nitrogen.
- Nutrient Management To Nutrient Stewardship
- N And P Responses For Oaten Hay
- Australia’s Declining Crop Yield Trends II: The Role Of Nitrogen Nutrition.
- Replacement P Can It Maintain Soil Fertility And Crop Production?
- Utilising Chicken Litter As A Crop Fertiliser – Do We Need To Apply Additional Conventional Fertiliser?
- Application Of Diffusive Gradients In Thin Films (DGT) To Measure Potassium And Sulphur Availability In Agricultural Soils
- Extractable Phosphorus (Colwell) Concentrations Of Soil After Banding Fertiliser With Seed In Relation To The Critical Phosphorus Requirement Of A Wheat Crop
- The Form And Fate Of Stubble Phosphorus In Cropping Soils
- Comparison Of Decision Tools To Improve The Nitrogen Management In Irrigated Maize Under Mediterranean Conditions In Spain
- Assessing Break Crop Options For Enhanced Soil Phosphorus Availability
- Effect Of Biochar On P Uptake From Two Acid Soils
- Ammonia Volatilisation Losses From Fertilisers Surface Applied To Vertosols In The Northern Australian Grains Region
- Pastures
- Seasonal Production Of Coloured Brome (Bromus Coloratus Steud) Cv. Exceltas, A New High Quality Perennial Temperate Pasture Grass
- Austrodanthonia Spp. In Grazed Native Pastures On The Monaro Tableland, NSW
- Southern Australian Feed Base Pasture Audit
- Use Of Non Fertiliser “growth Promotants” As Alternatives To Urea On High Production Pastures
- Pasture Legume Production Severely Reduced When Co Sown With Winter Forage Cereals
- Hard Seed Breakdown Rates Of Six Tropical Ley Pasture Legumes.
- Potential Of Desmanthus Spp. In Northern New South Wales
- Companion Legume Species Maximise Productivity Of Chicory (Cichoruim Intybus)
- Success Of Perennial Pasture Establishment At Different Sowing Times And Under A Cover Crop In The Mixed Farming Zone
- Summer Sowing: A New Alternative Technique To Introduce Annual Pasture Legumes Into Mixed Farming Systems
- Seed Rain Of Microlaena Stipoides
- Novel Cocksfoots For SE Australia – Establishment And Production
- A Survey Of Land Use And Management, North West Slopes Of New South Wales
- The Germination, Passage And Viability Of Desmanthus Virgatus (L.) Willenow Seed Through Sheep And Its Implication For Dispersal In Tropical Rangelands
- Low Input, High Quality Legume Hays For North Queensland.
- Evaluating New Phalaris Populations For Lower Rainfall Margins In SE Australia
- The Advantages And Disadvantages Of Pasture Cropping In Western Australia
- The Search For Sustainable Stocking Rates On Chinese Desert Steppe
- Improving Mineral Availability For Grazing Livestock In Australian Pasture Systems By Using Plantain And Lucerne
- A Survey Of Farmer Practice On The Establishment, Duration And Production Of Pastures In The Mixed Farming Zone Of Southern New South Wales
- To Under Sow Or Not? A Decision Support Tool To Determine The Most Profitable Method Of Pasture Establishment
- Nutritive Value Of Silage From Perennial Ryegrass, Plantain And Lucerne Pastures In South West Victoria
- Crop Yield, Pasture Yield, And Environmental Impact Of Pasture Cropping With Sub Tropical Perennials.
- Identifying Lucerne Growers’ Usage And Needs Through A Survey For Breeding New Generation Lucerne Varieties
- Opportunities For Grazing Crop/pasture Intercrops
- Weed Control Options For Pasture Cropping Systems In The West Midlands Of Western Australia
- Evaluating The Importance Of A Potential Source Of Error When Applying Shoot 15N Labelling Techniques To Legumes To Quantify The Below Ground Transfer Of Nitrogen To Other Species
- Important Principles Of Pasture Management Demonstrated Within The Cicerone Project’s Grazed Farmlet Experiment: Some Personal Reflections
- Tasmanian Pasture Resource Audit: Snapshot Of Functional Group Composition In 2011
- Increasing Soil Fertility And Altering Legume Grass Balance As Adaptation Strategies For Sustainable Livestock Production Under Climate Change
- Graziers, Pasture Seed Industry And Researchers Are Concerned About Pasture Productivity Decline.
- New Tall Fescue Cultivars For Medium To Low Rainfall Environments In Southern Australia
- Assessment Of Lotus Tenuis For Desirable Seed Production Characteristics
- Leucaena: Highlighting The Value Of Good Agronomy For Establishing Pasture Systems.
- EverFarm: Piloting A Strategic Decision Making Process To Improve Grazing Farm Systems.
- Pests
- Identifying A Non Destructive Technique To Assess Nematode Tolerance In Wheat Variety Trials.
- Have You Considered The Biosecurity Risks Of Your Agricultural Consultancy, Field Research Or Field Day?
- Susceptibility Of Annual Medics (Medicago Spp.) To Powdery Mildew (Erysiphe Trifolii)
- Transform™ Insecticide (sulfoxaflor) For Control Of Aphids In Canola
- Plenary Papers
- Chemical Free Agriculture: Obstacles And Assumptions
- What Has Australian Agronomy Contributed To World Food Security In The Last 20 Years, And What Lies Ahead?
- Are Australian Farmers, The Agricultural Research Community And Governments Capable Of Adapting To Changing Paradigms In Agronomy?
- Nutrient Supply And Management
- Thoughts On Capturing Opportunities And Overcoming Obstacles For Australian Agriculture Related To Electronic Agriculture
- Capturing Opportunities And Overcoming Obstacles For Australian Agronomy Related To Weed Management
- Raising The Genetic Potential Of Rainfed Crops Through Biotechnology Opportunities And Challenges.
- Precision Agriculture
- CliMate – A Smartphone App For Analysing Climate Data
- Field Based Rapid Phenotyping With Unmanned Aerial Vehicles (UAV)
- Leaf Nitrogen Determination Using Handheld Meters
- SoilMapp – Mobilising Australian Soil Data
- Controlled Traffic Experiences And Results In A Diversified Vegetable Industry
- The Integration And Validation Of Precision Management Tools In Mixed Farming Systems.
- Site Specific Measurements Of Apparent Electrical Conductivity (ECa) Correlate To Neutron Moisture Probe Counts: Towards The Spatial Measurement Of Soil Moisture Content For Precision Agriculture.
- Bridging The Yield Gap : Diagnosing Poor Performing Patches From Paddock To Farm Scale
- Improved Monitoring Of Soil Water Resources To Benefit Crop Management Decisions
- Electromagnetic Induction Methods For Monitoring Soil Water In Irrigated Cropping Systems.
- Low Induction Number EM Surveys For Assessing Land Management Attributes In Upland Environments.
- Detection Of Pasture Pests Using Proximal PA Sensors: A Preliminary Study Investigating The Relationship Between EM38, NDVI, Elevation And Redheaded Cockchafer In The Gippsland Region
- Use Of Mobile Devices In Extension And Agricultural Production A Case Study
- Examining The Potential For Active Optical Sensors To Provide Biomass Estimation In Improved And Native Pastures
- CropMate™ A Web Based Decision Support Tool Helping Farmers Make Agronomic Decisions Using Historic And Forecast Weather And Climate Data.
- Comparison Of Soil Conductivity Measured By ERT And EM38 Geophysical Methods Along Irrigated Paddock Transects On Black Vertosol Soils
- Mixing Grapes And Grain Scoping The Opportunity For Selective Harvesting In Cereals
- Soil Water Management
- Water Application Strategies For Lucerne
- Modelling Water Use Of Irrigated Forage Systems In Northern Victoria
- Using Soil Water Content To Predict Pasture Growth Rates
- APSoil Providing Soils Information To Consultants, Farmers And Researchers
- Management Strategies To Improve Water Use Efficiency Of Barley In Tasmania
- Potential Of Dual Gamma Ray And Electromagnetic Induction Sensors To Map Soil Type And Plant Available Water Capacity
- Yield Gap Analysis: Quantifying The Gap Between Farmers’ Wheat Yields And Water Limited Yield Potential
- Water Relations In Two Cultivars Of Napier Grass Under Variable Water Supply And Temperature Conditions
- Dynamic Deficits For Irrigated Cotton – Matching The Soil Water To Plant Requirements.
- Maintaining Yield / Reducing Nitrogen Losses In Water: A Sustainable Solution For Sugarcane Production Systems
- Water Use And Water Use Efficiency Of Wheat Under A Free Air CO2 Enrichment (FACE) Experiment
- Subsoil Manuring On Problem Clay Soils: Increasing Crop Yields To The Next Level
- Summer Fallow Management In 2010 Across Central West NSW
- Waterlogging, Anoxia And Wheat Growth In Surface Irrigated Soils
- Does Control Of Summer Fallow Weeds Improve Whole Farm Productivity In Both Mediterranean And Temperate Environments? A Simulation Analysis
- Utilising Soil Water Sensing And Modelling To Guide Grower Decision Making In Dryland Cropping Systems A Case Study
- Soil Water Express – A System To Generate Approximate Soil Water Characterisations And Current Soil Water Estimates From Minimal Input Data
- The Soil Evaporation Potentiality Index: A Toolkit Of Holistic Indicators To Monitor Soil Salinity And Associated Soil And Vegetation Degradation, For Agricultural Productivity In Upland Environments.
- Development, Growth And Water Extraction Of Seedling Lucerne Grown On Two Contrasting Soil Types
- Nitrous Oxide Emissions In Medium Rainfall Cropping Environments: Merely Interesting, Or Actually Important?
- Producing More With Less Using Retro Fit Telemetry To Reduce Energy And Water Consumption During Carrot Production
- LPAD: A Lysimetery Platform To Phenotype The Temporal Transpiration Pattern For Field Crops Over The Growing Cycle.
- Simulated Soil Water Balance Is Affected By Crop Rotation/irrigation Systems In The Irrigated Lockyer Valley, Queensland, Australia
- Climate Conditions Affected The Simulated Soil Water Balance Of An Irrigated Vegetable Rotation In The Lockyer Valley, Australia
- Utilising Chicken Litter As A Soil Improvement Agent
- Lucerne Creates A Dry Soil Buffer To At Least 5m
- Rice And Sweet Corn Response To High Levels Of Water Supply
- Semi Automated, Non Weighing, Pot In Bucket (PIB), Water Management In Pot Plant Culture.
- Micro Basin Technology In Ethiopia – Is It Worth The Effort?
- Can Changes In The Cropping/livestock Mix, Moisture Seeking Planting And Fallow Length Mitigate The Impacts Of Climate Change In Queensland?
- Agriculture And Languages Other Than English
- Water Extraction Of Solid And Skip Row Cotton
- Weeds
- Chemical Control Of Onion Grass (Romulea Rosea) In Native Pastures
- Annual Ryegrass (Lolium Rigidum) Control With Pre Emergent Herbicides Under Different Tillage Systems
- Novel Herbicide Tolerance In Lentils.
- Grain Yield Implications Of Crop Topping Pulses For Late Weed Control In South Eastern Australia
- Sustainable Weed Control In Cucurbit Crops: A Scoping Study
- Tolerance Of Trifolium Tumens (Talish Clover) To Pre And Post Emergent Herbicides
- Agriculture
- 1980 1st AAC
1 Sugarcane Research Institute, Yunnan Academy of Agricultural Sciences, Kaiyuan, Yunnan , China, 661600
Email: chenhuiysri@ yahoo.com
2 Sugarcane Research Institute ,Yunnan Academy of Agricultural Sciences, Kaiyuan, Yunnan ,China, 661600
Email fyhysri@sohu.com.
3 Sugarcane Research Institute ,Yunnan Academy of Agricultural Sciences, Kaiyuan, Yunnan ,China, 661600
Email caiqingysri@ sohu.com.
4 Laboratory of Cellular and Molecular Evolution, Kunming Institute of Zoology ,the Chinese Academy of Sciences,
Kunming ,Yunnan, China 650223 Email yapingzhang@mail.kiz.ac.cn
In this study, 195 accessions of Saccharum Spontaneum L. from different geographical populations in China were studied using random amplified polymorphic DNA (RAPD) analysis to estimate their genetic diversity. Phylogenetic analysis was done to assess relationships among haplotypes. A total of 266 bands were scored of which 145 were reproducibly polymorphic. The extent of genetic variability was between 62.95% and 37.05% within populations. The topology of the phylogenetic tree of S. spontaneum corresponds to the population geographical distribution. The genetic variation and diversity among different populations was consistent with existing knowledge on the geographical distribution of S. spontaneum in China. We tentatively propose that Yunnan province was the center of origin for S. spontaneum in China, based on previous collections and archeological information when combined with these results.
Based on our research on genetic diversity and phylogeny combined with previous collections and archeological information, Yunnan province is the center of origin for S. spontaneum in China. and
Key Words
Saccharum Spontaneum L., RAPD, genetic diversity, phylogeny
Introduction
China's vast territory, complicated geological formation and various climate types create favorable conditions for the evolution of wild sugarcane. Diverse sugarcane genetic resources evolved in these variable environments. Germplasm collections are an important component of sugarcane improvement programs as they provide breeders with sources of useful traits. Basic germplasm has often been used in sugarcane breeding. Most modern commercial sugarcane clones represent a complex interspecific combination between the genomes of S. officinarum and S. spontaneum (Besse et al. 1997). At present, sugarcane breeding in China is progressing towards the introduction of valuable genes, with positive effects on agronomically important traits, from new basic germplasm into modern sugarcane clones. This is especially true for genes from the species S. spontaneum, which has played a very important role in sugarcane breeding in the development of interspecific hybrids. S. spontaneum has the widest geographical distribution of the six species of the genus Saccharum (S. officinarum L., S. spontaneum L., S. barberi Jeswiet., S. sinense Roxb., S. robustum Brandes & Jeswiet ex Grassl. and S. edule Hassk.). S. spontaneum has many desirable characters including disease resistance, vigour, tillering ability, drought tolerance, water tolerance, frost tolerance, and general adaptability. Because of these traits, sugarcane breeders all over the world have considerable interest in the collection, maintenance, evaluation, and exploitation of its genetic potential (Tai et al. 1999). S. spontaneum clones are distributed throughout 8 provinces in China. Clones have been collected on a number of occasions, and maintained in the National Nursery for Sugarcane Germplasm Resources, Yunnan Sugarcane Research Institute, Kaiyuan, Yunnan province, China. In this paper, we examine the use of RAPDs for estimating the level of genetic diversity among 195 S. spontaneum accessions from different geographical populations in China, and discuss the implications of our results for the origin and diffusion of the wild sugarcane.
Materials and Methods
Materials
195 accessions from eight geographic populations were considered in this study (Table 1). All samples were obtained from the National Nursery for Sugarcane Germplasm Resources in Yunnan province, China.
Table 1. The geographical distribution and climatic characteristic in 8 geographical colonies of Saccharum spontaneum L in China.
Colony |
Number of samples |
Geographical distribution and climatic characteristic samples |
Yunnan (YN) |
86 |
locate in the southwest of China, attrib to semi-tropic, plateau, moisture and monsoon climate |
Sichuan(SC) |
36 |
locate in the southwest of China, attrib to semi-tropic, plateau, moisture and monsoon climate |
Guizhou(GZ) |
10 |
locate in the southwest of China, attrib to semi-tropic, plateau, moisture and monsoon climate |
Guangxi (GX) |
13 |
locate in the south of China, attrib to tropic, moisture and monsoon climate |
Guangdong(GD) |
18 |
locate in the south of China, attrib to tropic, moisture and monsoon climate |
Hainan(HN) |
14 |
locate in the south of China, attrib to tropic, moisture and monsoon climate |
Fujian(FJ) |
14 |
locate in the southeast of China, attrib to semi-tropic, moisture and monsoon climate |
Jiangxi(JX) |
4 |
locate in the southeast of China, attrib to semi-tropic, moisture and monsoon climate |
Methods
DNA was isolated from apical meristem and young leaf tissues using a modified method (Fan et al. 1999). PCR mixtures (10μL total) contained: 25ng DNA, 2.5mM dNTP, 0.2μM primers (products of Operon Technology Company),10 mM Tris-HCl (pH8.9), 50 mM KCl, 0.2mM BSA, 2.5mM MgCl2, and 1.0 unit of Taq polymerase (products of Takara Biotechnology Company). Reactions were overlaid with 20μL of mineral oil to prevent evaporation. Samples for enzymatic amplification were subjected to initial denaturation at 95°C for 3 min follow by 40 cycles of 94°C for 1min (denaturation), 36°C for 1min (annealing), and 72°C for 2min (extending) with a final extension at 72°C for 5min. Fragments generated by amplification were separated according to size electrophoresis on 1.5% agarose gel in 1×TAE, stained with ethidium bromide, and photographed by an EAGLE EYE imager (Williams et al. 1990; Wachiro et al. 1995; Chen et al. 2001).
Results
RAPD Analysis
Out of a total of 25 primers used, 20 (80%) generated polymorphic loci. A total of 266 bands were scored of which 145 (54.5%) were reproducibly polymorphic. The number of products generated by each primer varied from 7 to 15 with an average of 13. The size of the amplified fragments that were scored ranged from 0.2-2kb.
Genetic Diversity
The phenotypic frequencies detected with the 20 primers were calculated and used in estimating diversity (Ho) within population types (Table 2). The Jiangxi population exhibited the lowest within population variability (0.6937), but had the smallest population size, which affects Ho. The Yunnan population exhibited highest within population variability (1.7126).
Shannon’s index of phenotypic diversity was used to partition the diversity into within and between population components (Table 3). Primer OPI-08 detected the most within population variability, while primer OPA-19 detected the least.. An assessment of the proportion of diversity present within population Hpop/Hsp, compared with that between populations, (Hsp-Hpop)/Hsp, indicates that, on average, most of the diversity (62.95%) is detected between populations (38.15%-71.86%). 37.05% of the variation was detected within populations. However, the distribution of variability did vary slightly between and within populations with different primers.
Table 2. The genetic diversity index of 8 geographical colonies of Saccharum Spontaneum L. in China
Primer |
YN |
SC |
GZ |
GX |
GD |
HN |
FJ |
JX |
OPA-07 |
2.0387 |
1.7673 |
1.4536 |
1.3849 |
1.2767 |
1.0885 |
1.1456 |
0.9764 |
OPA-19 |
2.0909 |
1.8421 |
1.7654 |
1.4308 |
1.5692 |
1.2048 |
1.0923 |
0.8732 |
OPB-14 |
1.4758 |
1.8122 |
0.9091 |
0.7422 |
0.7288 |
0.7059 |
0.4378 |
0.6496 |
OPD-01 |
2.1492 |
1.9678 |
1.6216 |
1.4358 |
1.2697 |
0.9739 |
0.7262 |
0.6519 |
OPF-01 |
1.3591 |
1.2943 |
1.0249 |
0.9739 |
0.7618 |
0.7618 |
0.8732 |
0.6348 |
OPF-05 |
2.2241 |
1.9769 |
2.0349 |
1.3851 |
1.0624 |
1.0348 |
0.9792 |
0.7567 |
OPF-12 |
1.8526 |
1.9769 |
1.7743 |
1.3155 |
1.2349 |
0.9849 |
0.8492 |
0.6599 |
OPH-01 |
1.5101 |
1.3844 |
1.0989 |
0.9432 |
1.1143 |
0.8938 |
0.7607 |
0.8349 |
Primer |
YN |
SC |
GZ |
GX |
GD |
HN |
FJ |
JX |
OPH-19 |
1.8122 |
1.4278 |
1.3342 |
1.1893 |
0.9866 |
0.8654 |
0.7348 |
0.6549 |
OPI-08 |
1.4637 |
1.3859 |
1.2248 |
1.1049 |
1.0876 |
0.9703 |
0.8785 |
0.7786 |
OPJ-07 |
1.2917 |
1.0977 |
0.9771 |
0.8718 |
0.7185 |
0.6507 |
0.5873 |
0.4336 |
OPJ-09 |
1.7298 |
1.6348 |
1.3285 |
1.2853 |
1.0928 |
0.9385 |
0.6418 |
0.5186 |
OPJ-14 |
1.2875 |
1.4637 |
1.1524 |
1.0335 |
0.8785 |
0.8324 |
0.6545 |
0.5448 |
OPJ-18 |
1.8819 |
1.6405 |
1.4969 |
1.1721 |
0.9898 |
0.8671 |
0.7703 |
0.6479 |
OPK-18 |
1.9416 |
1.8803 |
1.6581 |
1.3272 |
1.2717 |
1.0068 |
0.8781 |
0.7589 |
OPL-17 |
1.9769 |
1.7734 |
1.5266 |
1.6534 |
1.2548 |
0.9849 |
0.8834 |
0.6599 |
OPM-04 |
1.6309 |
1.3154 |
1.4403 |
1.1191 |
1.0348 |
0.8779 |
0.5946 |
0.6343 |
OPM-07 |
1.7898 |
1.9416 |
1.6034 |
1.4432 |
1.0781 |
1.2249 |
0.9789 |
0.8744 |
OPN-02 |
1.1143 |
1.5533 |
0.9073 |
0.6471 |
0.8411 |
0.5489 |
0.5302 |
0.4812 |
OPN-11 |
1.6309 |
1.2951 |
1.4038 |
1.1438 |
0.9779 |
1.0433 |
0.9342 |
0.8495 |
Average |
1.7126 |
1.6216 |
1.3868 |
1.1801 |
1.0615 |
0.9230 |
0.7965 |
0.6937 |
Table 3. Partitioning of the genetic diversity into within and between populations for 20 primers
Primer |
Hsp |
Hpop |
Hpop/Hsp |
(Hsp-Hpop)/Hsp |
OPA-07 |
3.9898 |
1.3915 |
0.3487 |
0.6513 |
OPA-19 |
5.2715 |
1.4836 |
0.2814 |
0.7186 |
OPB-14 |
2.3744 |
0.9327 |
0.3928 |
0.6072 |
OPD-01 |
4.0458 |
1.3495 |
0.3335 |
0.6665 |
OPF-01 |
1.6233 |
0.9605 |
0.5917 |
0.4083 |
OPF-05 |
4.8902 |
1.4318 |
0.2928 |
0.7072 |
OPF-12 |
3.9773 |
1.3311 |
0.3347 |
0.6653 |
OPH-01 |
1.9903 |
1.0675 |
0.5364 |
0.4636 |
OPH-19 |
2.8188 |
1.1256 |
0.3993 |
0.6007 |
OPI-08 |
1.7977 |
1.1118 |
0.6184 |
0.3815 |
OPJ-07 |
1.7671 |
0.8285 |
0.4688 |
0.5312 |
OPJ-09 |
3.2252 |
1.1463 |
0.3554 |
0.6446 |
OPJ-14 |
2.2187 |
0.9809 |
0.4421 |
0.5579 |
OPJ-18 |
3.4079 |
1.1833 |
0.3472 |
0.6528 |
OPK-18 |
4.3257 |
1.3403 |
0.3098 |
0.6902 |
OPL-17 |
4.5241 |
1.3392 |
0.2961 |
0.7039 |
OPM-04 |
2.5573 |
1.0809 |
0.4227 |
0.5773 |
OPM-07 |
4.0844 |
1.3668 |
0.3346 |
0.6654 |
OPN-02 |
2.3378 |
0.8279 |
0.3541 |
0.6459 |
OPN-11 |
2.0307 |
1.1598 |
0.5711 |
0.4289 |
Average |
3.1619 |
1.1719 |
0.3705 |
0.6295 |
Genetic Differentiation and Genetic Relationship
To examine the genetic differentiation and genetic relationship between the different populations, a genetic distance matrix based on the proportion of shared fragments (Nei 1978) was used to establish the level of relatedness between the different populations of S. spontaneum studied. Figure 1 shows a dendrogram generated by UPGMA cluster analysis, based on the estimates of genetic distances between populations.
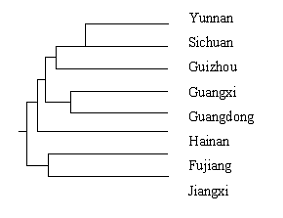
Figure 1. A dendrogram generated by UPGMA cluster analysis, based on the estimates of genetic distances between populations.
Conclusion
This study demonstrated that S. spontaneum clones within the Yunnan population exhibited more extensive genetic variability, as well as more abundant genetic diversity and many original types of wild sugarcane. Yunnan province has the widest geographical distribution of S. spontaneum of all provinces in China. Moreover, according to our results, studies on sugarcane collection, maintenance, evaluation and exploitation of its genetic potential over many years, and related archeological information, we tentatively proposed that Yunnan province was the possible origin center of S. spontaneum in China, which then spread into Sichuan, Guizhou and Guangxi provinces. S. spontaneum in Guangxi further spread into Guangdong province, followed by spread into Fujiang and Jiangxi provinces northward and Hainan province southward.(Figure 2)
Figure 2. The possible origin center and diffusion paths of Saccharum Spontaneum L. in China were demonstrated by the dot and the arrows respectively.
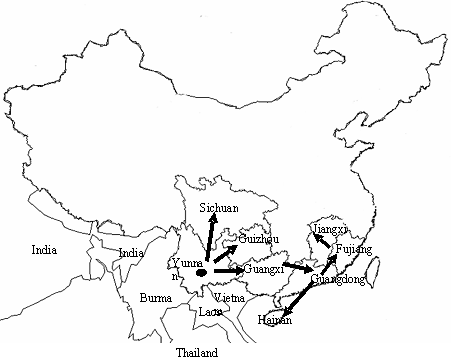
References
Besse P, McIntyre CL and Berding N (1997). Characterisation of Erianthus Sect. Ripidium and Saccharum Germplasm(Andropogoneae-Saccharinae) using RFLP Markers. Euphytica 93, 283-292.
Chen H, Fan YH, Shi XW, Cai Q, Zhang M and Zhang YP (2001). Research of Genetic Diversity and Systemic Evolution in Saccharum Spontaneum L. Crop Transaction 5, 645-652.
Fan YH, Cai, Q, Su B and Zheng YP (1999). Effects of DNA extraction and purification processes on the RAPD results of six Saccharinae species. Southwest China Journal of Agricultural Sciences 1, 1-7.
Nei M (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89, 583-590.
Tai PYP, Miller JD and Legendre BL (1999). Preservation of Saccharum spontaneum germplasm in the world collection of sugarcane and related grasses through storage of true seed. Sugar Cane 3, 4-10.
Wachira FN, Waugh R, Hackett CA and Powell W (1995). Detection of genetic diversity in tea (Camellia sinensis) using RAPD markers. Genome 38, 201-210.
Williams JG, Kubelik AR, Livak KJ, Rafalski JA and Tingey SV (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research 22, 6531-6535.




