1Wagga Wagga Agricultural Institute, PMB, Wagga Wagga, NSW 2650; Australia, 2Research Institute for Bioresources, Okayama University, Kurashiki, 710-0046, JAPAN.
Introduction
Acid soils limit the cultivation of barley (Hordeum vulgare L.) in parts of NSW, Victoria and Western Australia with over 450-500 mm rainfall. Most of the cultivated barley grown in Australia are sensitive to soil acidity and therefore, yield potential of these cultivars cannot be achieved on acid soils. To neutralise the soil acidity, application of lime is being used extensively in some areas. It has been observed that lime tends to move slowly into the sub-surface soil and tolerant cultivars perform better even after lime application on soils where acidity is deeper than the incorporation of lime (Scott et al., 1997). Breeding for acid soil tolerance is one of the important objectives of the barley improvement program in Australia.. As a result, two cultivars tolerant to soil acidity, Brindabella and Yambla have been released by Wagga Wagga Agricultural Institute. Selection of tolerant genotypes for soil acidity (aluminium, Al) using the haematoxylin stain test is very unsatisfactory. The other methods like screening in solution culture (Moroni et al 1999) is labour intensive, time consuming, and can handle only small populations. Hence, there is a need to develop a rapid screening method to select for aluminium tolerance. Molecular markers are highly efficient as they improve the efficiency of conventional plant breeding by carrying-out selection not directly on the trait of interest but on molecular markers linked to that trait (Mohan et al 1997). The recently developed technique of amplified fragment analysis has the potential to detect higher numbers of polymorphic loci than other PCR based techniques (Thomas et al 1995, Raman et al 1999). Here we report a DNA marker closely linked with tolerance to aluminium using AFLP technique (Vos et al 1995) on bulked segregant pools (Michelmore et al 1991) derived from F2 progeny of Yambla (WB 220) x WB 229.
Material and Methods
Screening of genotypes for aluminium tolerance:
Two lines of barley WB220 (Yambla) and WB229 were screened for aluminium tolerance using an acid soil (pH 4.13) from near Binnaway, NSW and in a hydroponic bio-assay developed at WWAI (Moroni et al 1999). These two genotypes and their 67 F2s were tested in a nutrient solution with added Al. The seeds of the parental lines and their derivatives were surface sterilised with 1% sodium hypochlorite and germinated in petri dishes for seven days. Thereafter, the plants were further grown in nutrient solution (pH 4.0) with added aluminium. The pH of nutrient solution culture was maintained at 4.3 throughout the experimentation. The initial Al concentration in solution was 100μM and this damaged root growth in all genotypes. The concentration was lowered to 50μM after 4 days. Plants were scored by three of us after a further 7 days in this solution for growth and regrowth of both the seminal and lateral roots. We identified 17 F2s where we all scored regrowth on seminals and laterals, and 17 genotypes where we all failed to find regrowth in any root. The remaining 33 genotypes had regrowth in some roots or our scores were not in agreement. In a second test of F3 genotypes derived from the tolerant and sensitive F2s a similar method was used but with 50 μM initially with a lowering to 10 μM. Only seminal roots were scored for regrowth.
DNA Extraction:
The leaves (8-10 cm) were collected from 21 day old seedlings and were used for DNA extraction in 2 ml round bottom Eppendorf tubes. The tubes containing tissues were frozen in liquid nitrogen and ground into powder with mini-pestle and mortar (Eppendorf). The DNA extraction was carried-out as described by Guidet et al (1991).
Bulk Segregant Analysis:
The DNA of 15 F2 plants each of tolerant and sensitive reaction to aluminium were pooled and the bulk segregant analysis was performed (Michelmore et al 1991) using AFLP analysis (EcoR1/Mse1). The polymorphic loci were further analysed at single plant level.
AFLP Analysis:
This was performed as described by Vos et al (1995) by using Large Genome AFLP System (Gibco-BRL, USA). About 200 nanogram of DNA was digested with EcoR1 and Mse1 enzymes and digested fragments were ligated with adaptors. The ligated DNA fragments were amplified in 20 μl volume by using 33P labelled different E- and unlabelled M- primers. The amplified products were equally mixed with formamide loading buffer (98% formamide, 10mM EDTA pH8.0, 0.5% bromophenol blue and 0.5% xyanol cyanol (w/v) and were denatured at 900C for 3 min. Electrophoresis was carried out at 120 W at 50 0C for 2 hrs using 5 per cent polyacrylamide gels (0.4 mm) containing 7.5M urea. The denatured PCR products (3μl) were loaded on the gels. After electrophoresis, the gels were transferred on Whatman 3mm filter papers and dried on gel drier (Bio-Rad). The dried gels were used further for autoradiography.
Linkage Analysis:
This was performed using MAPMAKER/EXP 3.0 (Lincoln et al 1993).
Results
Nutrient solution enabled the selection of tolerant, sensitive and segregating genotypes to aluminium stress. Both the parental lines were regarded as tolerant to aluminium based on field experience and they were not differentiated in the Binnaway soil. In the nutrient solution experiment Yambla was found to be more tolerant under high Al stress (Moroni et al. 1999) (Table 1).
Likewise, differential root growth was observed under high Al stress between tolerant and sensitive individuals of the F2 progeny when grown in solution culture. Among 67 F2 plants of Yambla x WB229, 17 each of tolerant and sensitive plants and 33 segregants (intermediates) were found which fits 1:2:1 Mendelian ratio. This indicates that aluminium tolerance in Yambla/WB229 is controlled by single gene. Among 32 primer combinations, 98 polymorphic loci were found. Two loci were found to be closely linked with gene governing aluminium tolerance. The allele amplified with primer E-ACG/M-CAA was more closer and seems to be linked with tolerance to aluminium (Fig 1).
Table 1. Differential Tolerance of Yambla and WB229 to Aluminium

*Significant different at p = 0.05 according to Duncan’s multiple range test
# Weights are expressed as a percentage of control.
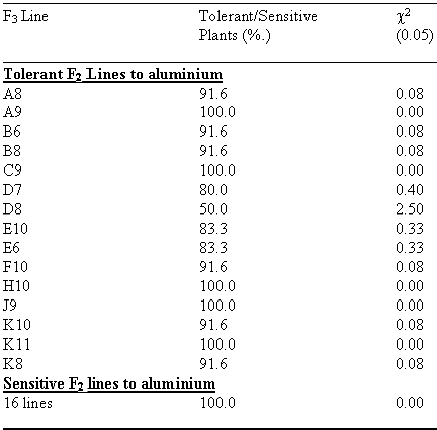
For the phenotypic expression of tolerance, the progeny testing using F3 lines of the selected F2s was done using solution culture (Table 2).
The results were found to be quite consistent, as most of the F3 plants have shown high frequency of consistency of being tolerant or sensitive to aluminium in F2. The F2 lines didn’t show any significant segregations in F3 generation (χ2 = 0.006, P> 0.995). Attempts are being made to map the aluminium tolerance on a molecular map of barley.

Figure 1. Bulk Segregant Analysis for Tolerance to Aluminium in Yambla x WB229. BT and BS: bulk DNA of Tolerants and Sensitives respectively, and 1-22: F2 plants of Yambla x WB229 {1-12: Tolerants and 13 to 22: Sensitives.
Acknowledgments
The authors are thankful to NSW Agriculture and the Acid Soil Action Initiative and the Grains Research and Development Corporation, Australia to provide financial support in project.
Table 2. Progeny Testing of F2 derivative in Solution Culture
References
1. Lincoln, S. E., Daly, M. J. and Lander, E. S. (1993). Research Technical Report, 3rd Edition p 1-95
2. Michelmore, R. W., Para, I., and Kasseri, R. V. (1991). Proc. Natl. Acad. Sci., USA, 88: 9828-9832.
3. Moroni, J. S., Scott, B. J., Sato, K., Read, B. J., and Raman, H. (1999). 11th Australian Plant Breeding Conference. Adelaide, Vol 2: 143.
4. Mohan, M, Nair, S., Bhagwat, A., Krishna, T. G., Yano, M., Bhatia, C. R., and Saski, T. (1997). Mol. Breed. 3: 87-103.
5. Thomas, C. M., Vos, P., Zabeau, M., Jones, D. A., Norcott, K. A., Chadwick, B. P., and Jones, J. D. G. (1995). The Plant J. 8(5): 785-794.
6. Raman, H., Sato, K., and Read B. J. (1999). Proc. 9th Australian Barley Technical Symposium, 12-16 September, Melbourne, Australia (this vol).
7. Scott, B. J., Conyers, M. K., Poile, G. J., and Cullis, B. R. (1997). Australian Journal of Agricultural Research 48, 843-854.
8. Vos, P., Hogers, R., Bleeker, M., Reijans, M., van de Lee, T., Hornes, M., Freiters, A., Pot, J., Peleman, J., Kuiper, M., Zabeau, M., 1995. Nucleic Acid Res. 23: 4407-4414.





